![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 3291_5903_10007_H94MGADXX_V_CF71_ATCACG_R2.fastq |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 7948472 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 38 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
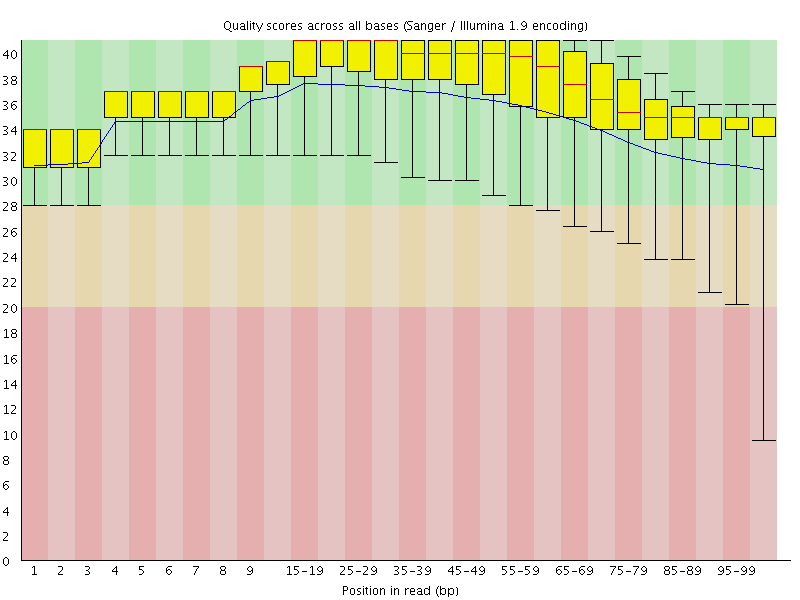
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
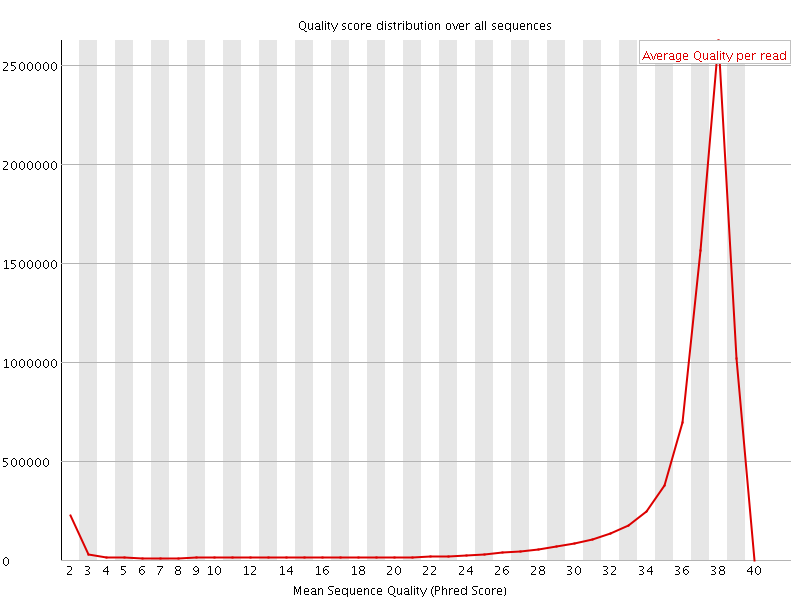
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
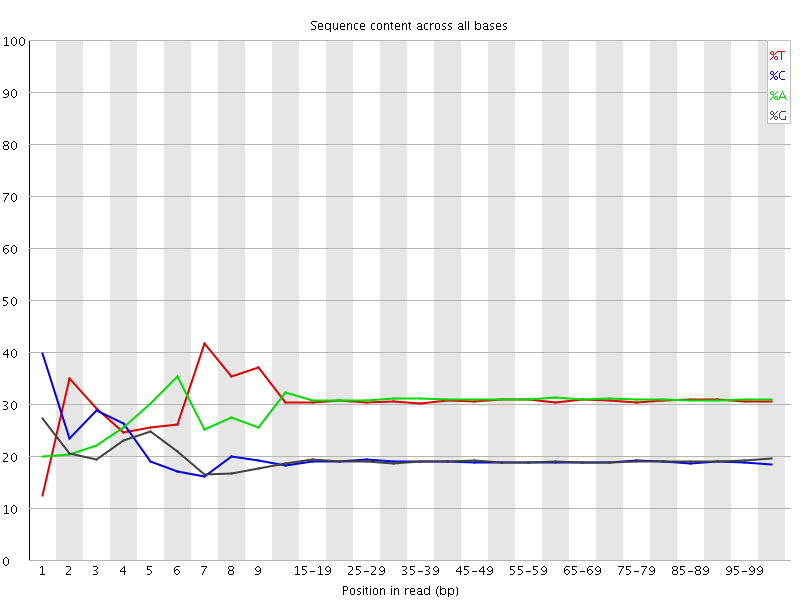
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
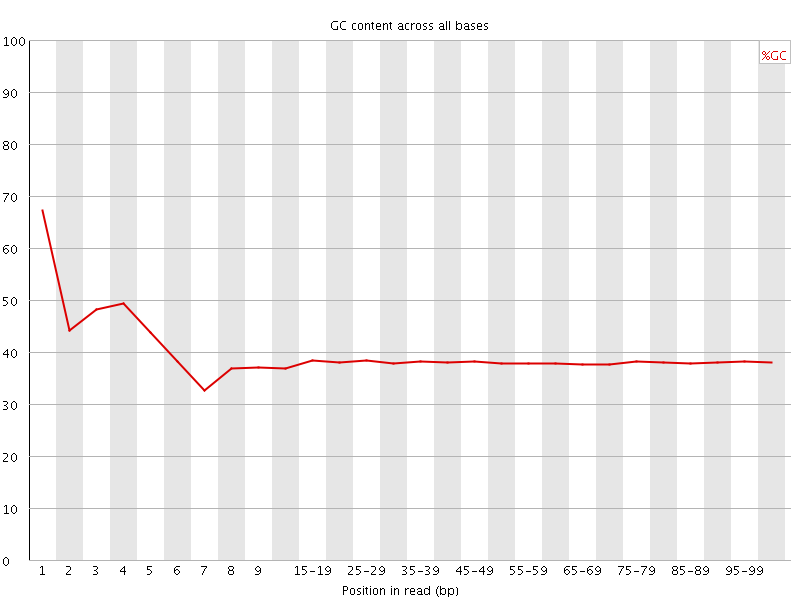
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
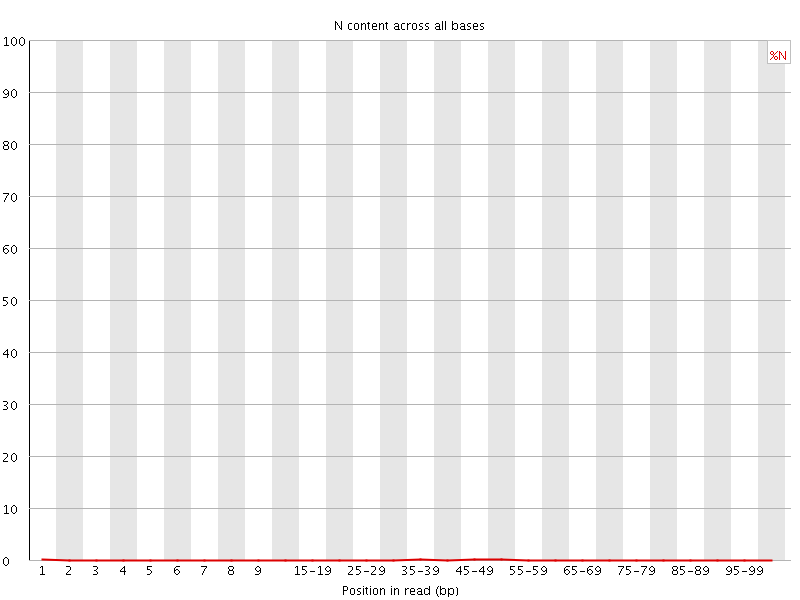
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
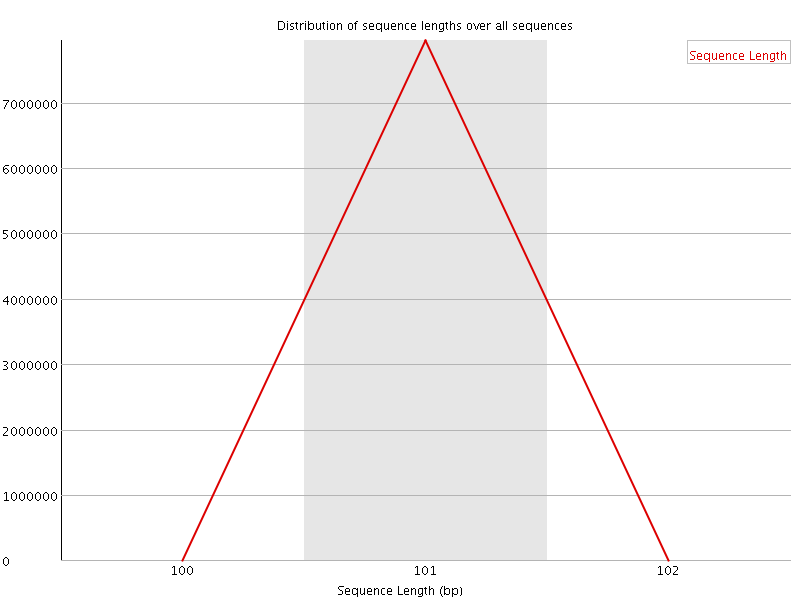
![[FAIL]](Icons/error.png) Sequence Duplication Levels
Sequence Duplication Levels
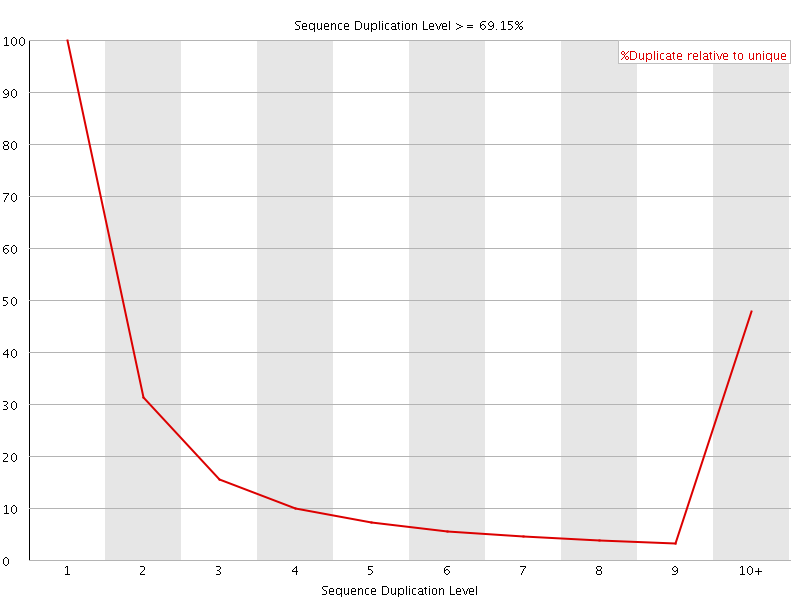
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GCCCCAACCAAAGTTATCTTATTAACTAACTAAATATAAATTATTAAAGC | 11609 | 0.14605322884700356 | No Hit |
| CCCCAACCAAAGTTATCTTATTAACTAACTAAATATAAATTATTAAAGCT | 8572 | 0.10784462724407912 | No Hit |
| CTCCTGTTCACTTTTCTTCCCCAATTCGAGGCTCTACACGCAAATGGCTG | 7990 | 0.10052246519834253 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
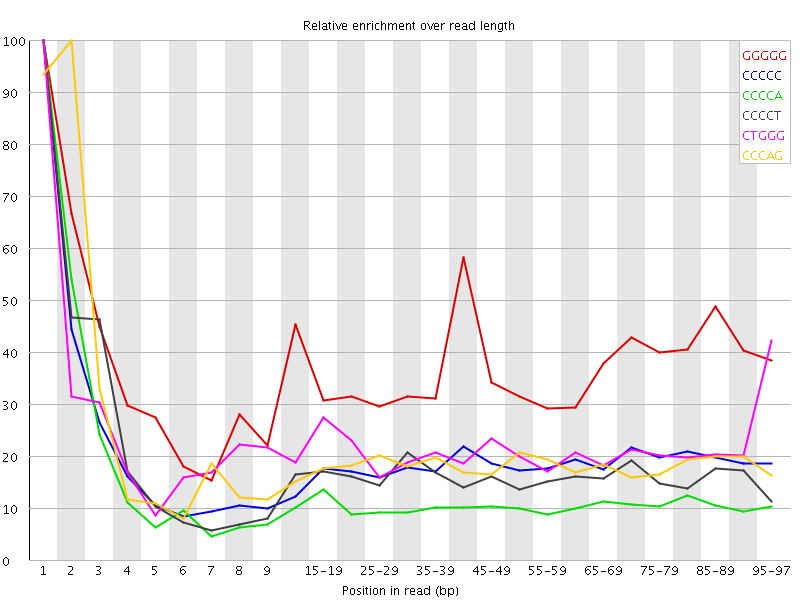
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GGGGG | 626790 | 3.0825377 | 8.206619 | 1 |
| CCCCC | 585620 | 2.7630394 | 14.415421 | 1 |
| CCCCA | 605790 | 1.808612 | 15.337283 | 1 |
| CCCCT | 580185 | 1.7328681 | 10.063681 | 1 |
| CTGGG | 565845 | 1.7326288 | 7.904476 | 1 |
| CCCAG | 533480 | 1.6059961 | 8.16166 | 2 |
| GCCCC | 328125 | 1.5610378 | 12.932515 | 1 |
| GCCCA | 512145 | 1.5417688 | 6.2249 | 1 |
| CCTGC | 511635 | 1.5408566 | 6.0753517 | 1 |
| CCTCC | 509190 | 1.5208237 | 6.577645 | 1 |
| GGCCC | 314140 | 1.5069556 | 8.290693 | 9 |
| CCCAA | 794295 | 1.5005739 | 6.2748036 | 2 |
| CTGCA | 778240 | 1.4830906 | 5.267369 | 1 |
| CTGGC | 487850 | 1.4814647 | 5.4757714 | 1 |
| TGGCC | 483040 | 1.4668584 | 6.411478 | 8 |
| CCTGG | 481475 | 1.4621059 | 5.7441053 | 1 |
| CCCTG | 477640 | 1.4384762 | 6.600274 | 1 |
| CTCCC | 481610 | 1.4384489 | 9.444488 | 1 |
| CGGGG | 281895 | 1.3748986 | 7.3834233 | 1 |
| CTCCT | 721430 | 1.3640207 | 8.082664 | 1 |
| CTCCA | 694665 | 1.3128846 | 6.1432767 | 1 |
| CCAGT | 686670 | 1.308586 | 6.6544814 | 3 |
| TCCTG | 659140 | 1.2566301 | 6.0346246 | 2 |
| CCGGG | 258880 | 1.2522147 | 5.814727 | 1 |
| CCCGG | 259670 | 1.2456586 | 6.977783 | 1 |
| CCCCG | 261335 | 1.2432882 | 9.964437 | 1 |
| CTGGA | 634435 | 1.2191141 | 5.5151777 | 2 |
| CCTGT | 618025 | 1.1782457 | 5.867778 | 3 |
| CGCCC | 244625 | 1.1637908 | 5.4118385 | 1 |
| CCTCG | 364410 | 1.097469 | 6.344393 | 1 |
| GTCCA | 550995 | 1.0500304 | 5.2497897 | 1 |
| CTCGG | 331950 | 1.0080398 | 7.759554 | 1 |
| GGATG | 519845 | 1.0072427 | 5.021434 | 4 |
| GTCCT | 497525 | 0.9485161 | 5.7184205 | 1 |
| CTCCG | 299725 | 0.90266156 | 6.75234 | 1 |
| CCCGC | 189215 | 0.9001809 | 6.409669 | 1 |