![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 3291_5903_10008_H94MGADXX_V_CF34_CGATGT_R1.fastq |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 9752597 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
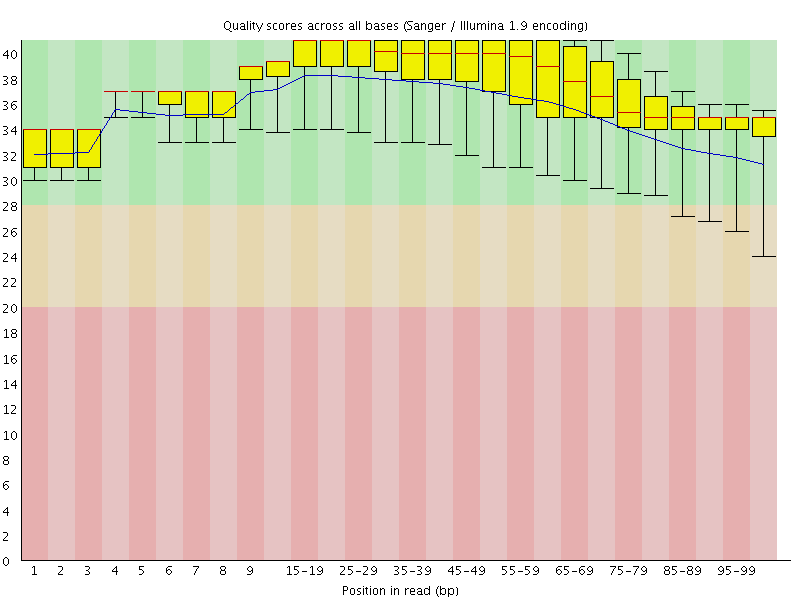
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
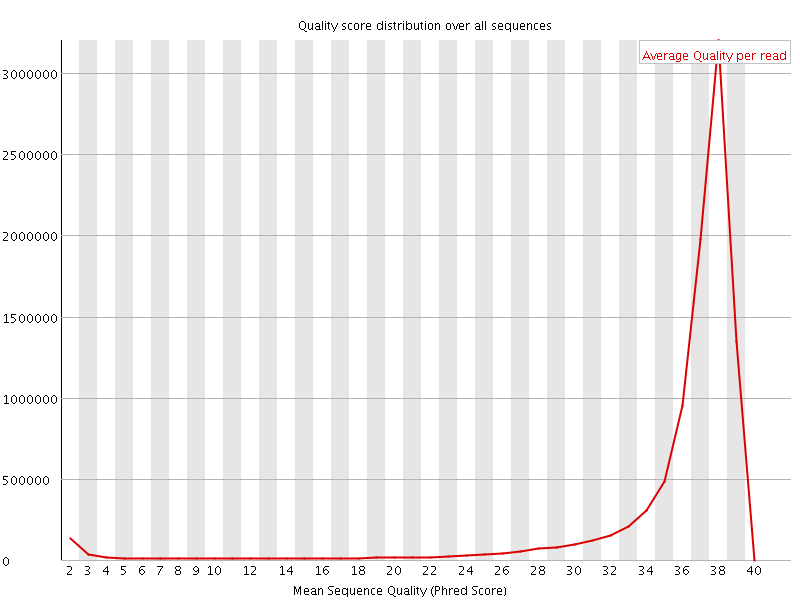
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
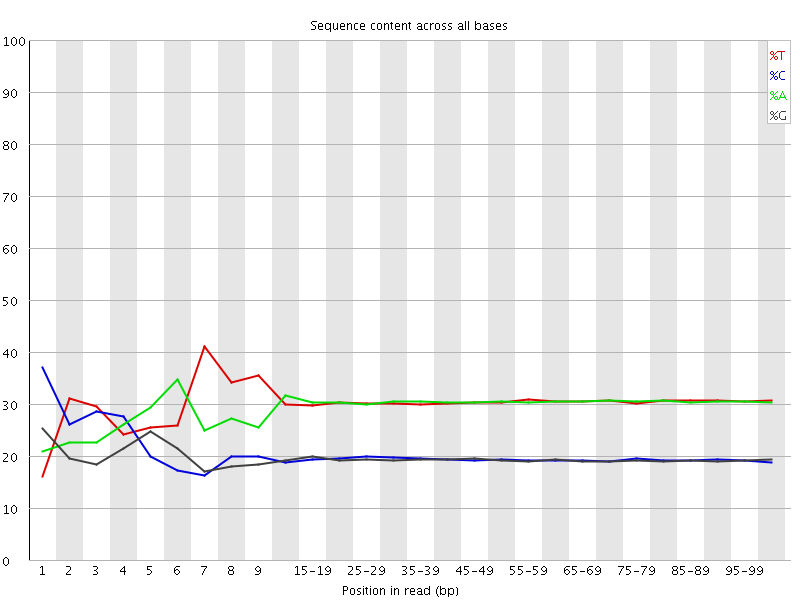
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
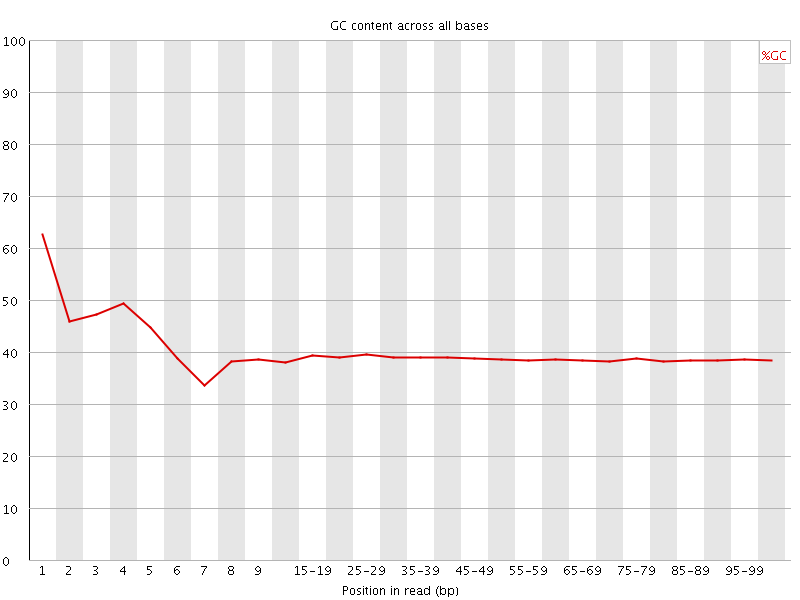
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
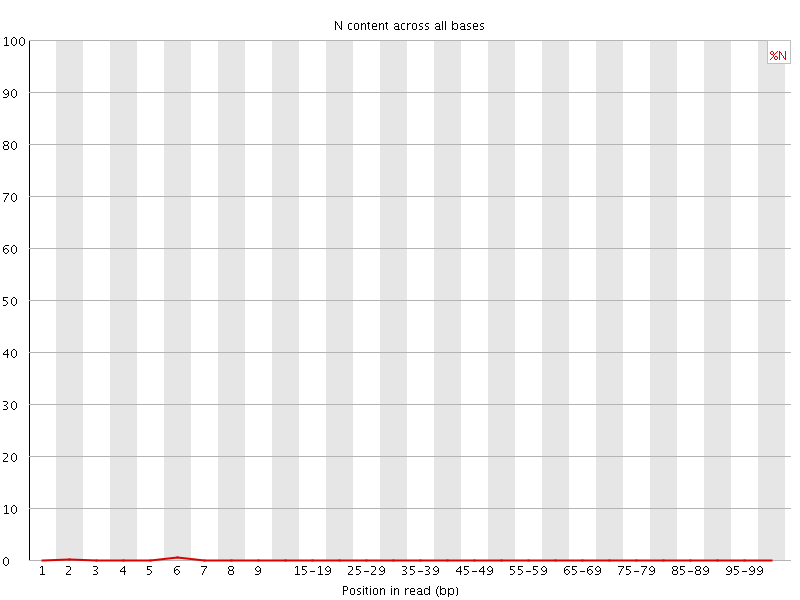
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
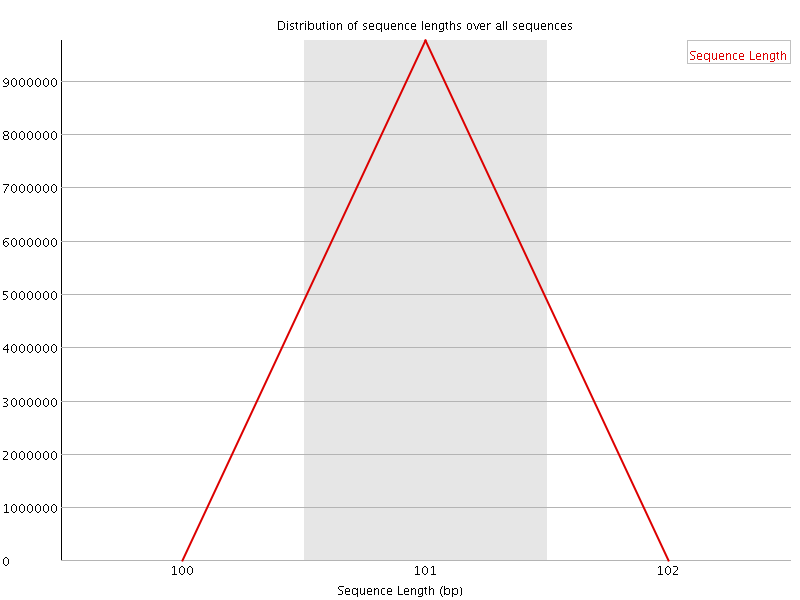
![[FAIL]](Icons/error.png) Sequence Duplication Levels
Sequence Duplication Levels
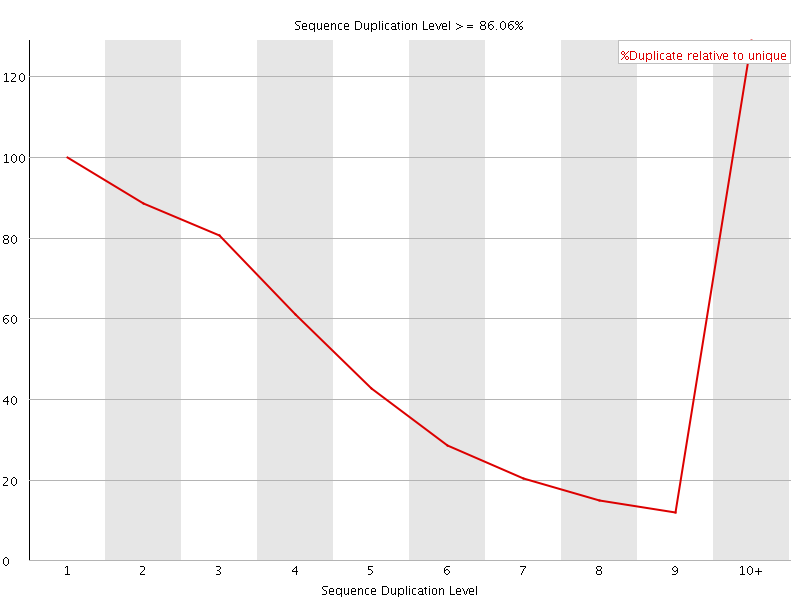
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| ACTAGTATGGCCCGGGGGATCCTACGTTCCAAATGCAGCGAGCTCGTATA | 28097 | 0.28809762158735774 | No Hit |
| ACTGGATGCATCTGCAGGATATCGCGGCCGCTCGACGTAGAACTCAATCT | 25507 | 0.2615405927262246 | No Hit |
| GATCCTTATCTGTCAAAACCGCTAATGTCCGTTCTAAGACCGTCTGGAGA | 24623 | 0.2524763404045097 | No Hit |
| ACTAGTATGGCCCGGGGGATCCTTATCTGTCAAAACCGCTAATGTCCGTT | 20542 | 0.2106310760098054 | No Hit |
| TCTGCAGGATATCGCGGCCGCCATCTGCCCTACGTTTGAGGGTTATAAGC | 19098 | 0.1958247633937914 | No Hit |
| ACTGGATGCATCTGCAGGATATCGCGGCCGCCATCTGCCCTACGTTTGAG | 18493 | 0.18962128754012905 | No Hit |
| CTCCTGTTCACTTTTCTTCCCCAATTCGAGGCTCTACACGCAAATGGCTG | 17594 | 0.1804032300319597 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
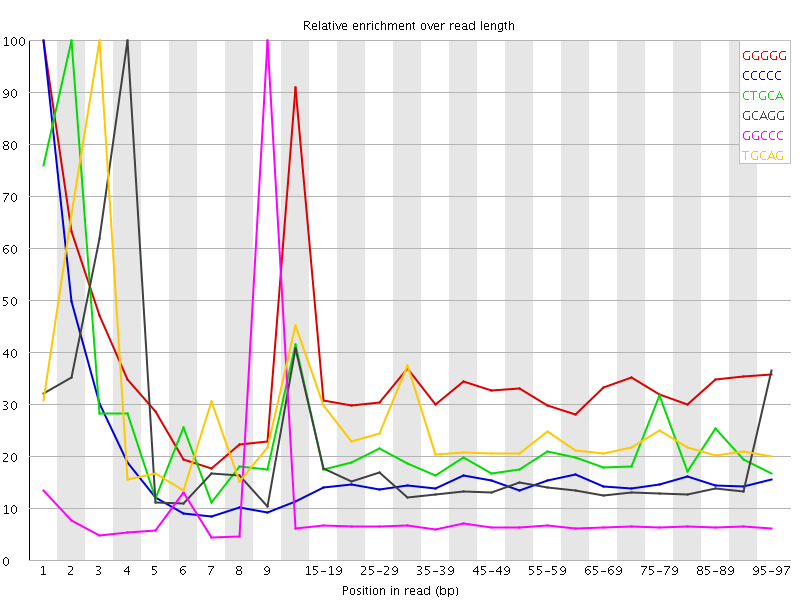
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GGGGG | 745270 | 2.8104672 | 7.7699733 | 1 |
| CCCCC | 608260 | 2.0847485 | 13.173693 | 1 |
| CTGCA | 1165820 | 1.7453802 | 7.803006 | 2 |
| GCAGG | 724645 | 1.7259095 | 9.699956 | 4 |
| GGCCC | 480075 | 1.7095181 | 22.517588 | 9 |
| TGCAG | 1117595 | 1.7054662 | 6.694987 | 3 |
| CCCCA | 721290 | 1.6221921 | 10.841608 | 1 |
| TGGCC | 685540 | 1.5942758 | 15.6607065 | 8 |
| GGCCG | 433000 | 1.5716381 | 5.5508776 | 25-29 |
| GCCCA | 670460 | 1.53697 | 5.74557 | 1 |
| CGGCC | 430040 | 1.5313464 | 5.44718 | 25-29 |
| TCTGC | 1010825 | 1.5061654 | 6.1648192 | 1 |
| CCTGG | 642350 | 1.4938343 | 5.15015 | 1 |
| CCCAG | 651540 | 1.4935976 | 6.7254605 | 1 |
| CCTCC | 663375 | 1.4848739 | 6.169593 | 1 |
| CCCAA | 1004505 | 1.4824243 | 5.3233185 | 2 |
| CCCTG | 637130 | 1.4536458 | 6.8589196 | 1 |
| CTGGA | 952205 | 1.4530786 | 10.816375 | 2 |
| ACTGG | 925935 | 1.4129902 | 9.480905 | 1 |
| CAGGA | 918890 | 1.4089129 | 6.1767874 | 5 |
| ATGGC | 916785 | 1.3990272 | 10.434405 | 7 |
| CGGGG | 374085 | 1.3839968 | 5.895139 | 10-14 |
| CCAGT | 919280 | 1.3762785 | 5.636821 | 3 |
| GCCCC | 393635 | 1.3751758 | 7.655118 | 1 |
| CCCCT | 614265 | 1.3749479 | 9.563521 | 1 |
| GCCGC | 384955 | 1.3708014 | 5.1961594 | 25-29 |
| CTCCA | 929970 | 1.3659267 | 5.4162126 | 1 |
| TGCAT | 1396665 | 1.3655792 | 6.7993493 | 7 |
| ATGCA | 1376395 | 1.352165 | 7.5492616 | 6 |
| CTCCC | 569650 | 1.2750833 | 8.459787 | 1 |
| CCTGT | 825580 | 1.2301435 | 7.3892074 | 3 |
| CTCCT | 819420 | 1.1978518 | 8.670532 | 1 |
| CCGGG | 329880 | 1.1973487 | 5.1415524 | 10-14 |
| CCCGG | 334970 | 1.1928079 | 5.506655 | 1 |
| TCCTG | 795615 | 1.1854947 | 6.5780582 | 2 |
| GATGC | 776365 | 1.1847442 | 9.780163 | 5 |
| GCATC | 777330 | 1.1637616 | 9.029988 | 8 |
| CATCT | 1179495 | 1.1314117 | 6.2666473 | 9 |
| GGATG | 700280 | 1.0892572 | 10.225376 | 4 |
| GCCCG | 304475 | 1.0842171 | 5.0683937 | 10-14 |
| CCCCG | 308485 | 1.0777017 | 7.656816 | 1 |
| CCTCG | 436625 | 0.99618304 | 5.341126 | 1 |
| TGGAT | 963890 | 0.96062124 | 7.2262664 | 3 |
| GATCC | 629515 | 0.9424637 | 7.5723352 | 1 |
| TATGG | 807735 | 0.80499583 | 6.4280624 | 6 |
| GTATG | 796780 | 0.794078 | 6.6271405 | 5 |
| CTCCG | 340940 | 0.7778726 | 5.0437803 | 1 |
| CTAGT | 543745 | 0.5316428 | 6.3062143 | 2 |
| ACTAG | 526980 | 0.51770306 | 6.161765 | 1 |