![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 3291_5903_10008_H94MGADXX_V_CF34_CGATGT_R2.fastq |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 9752597 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
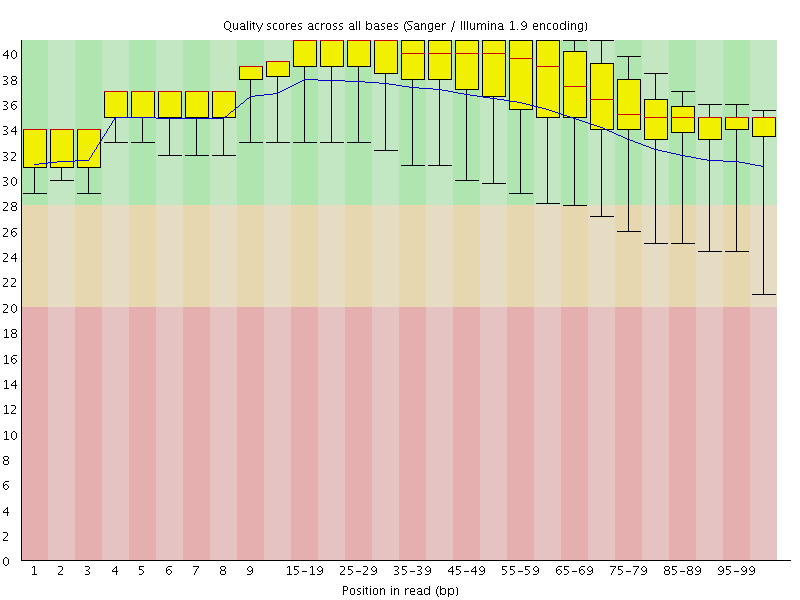
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
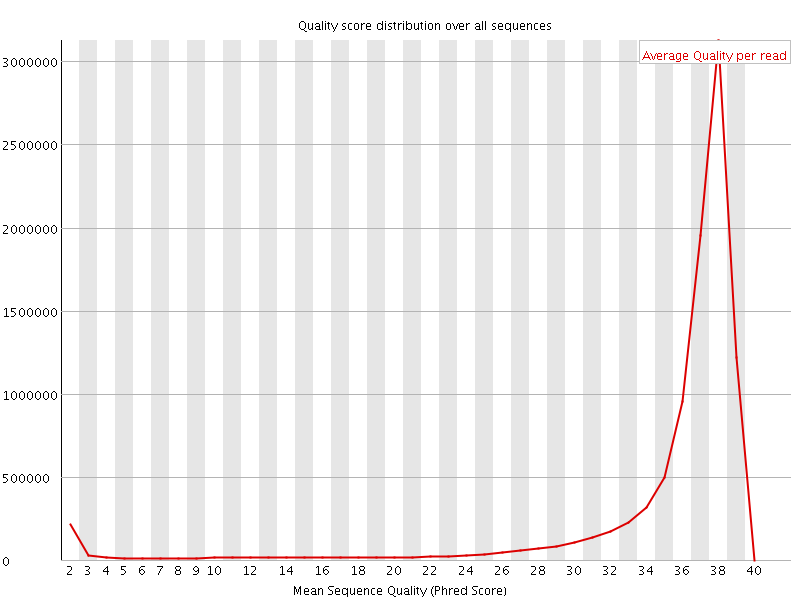
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
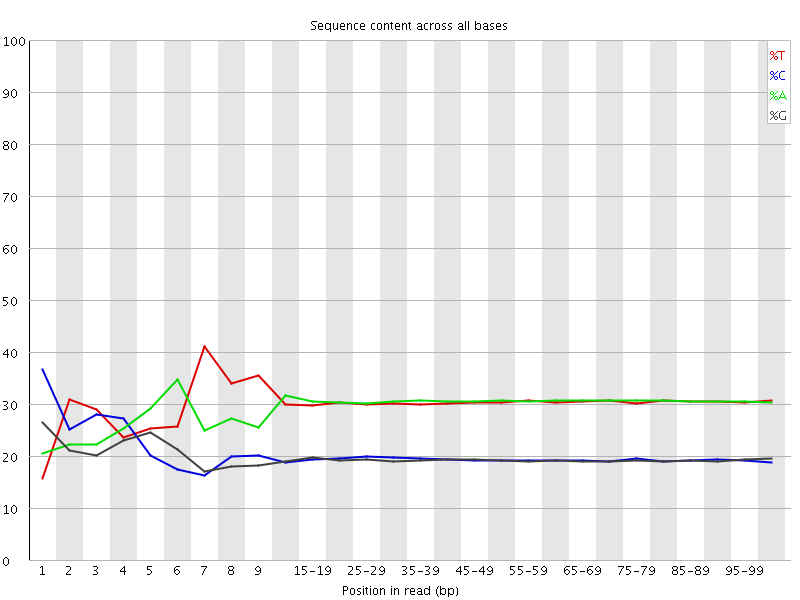
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
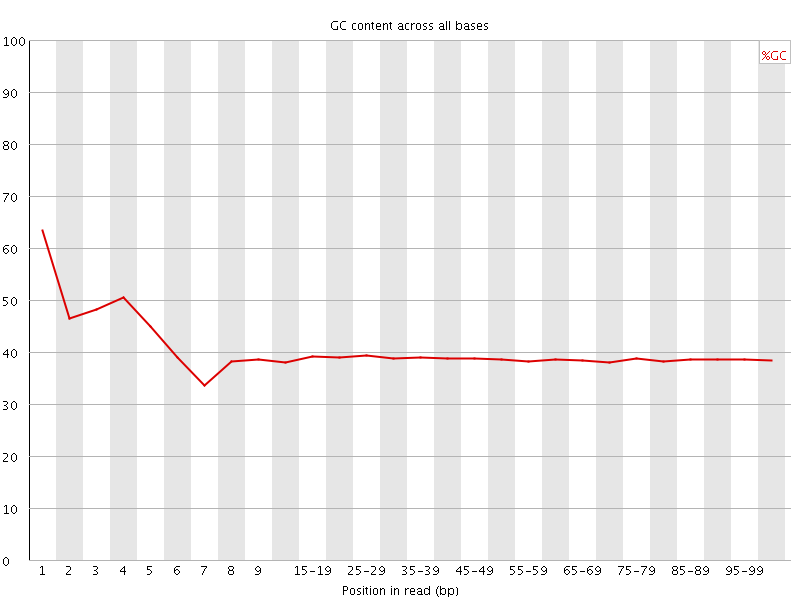
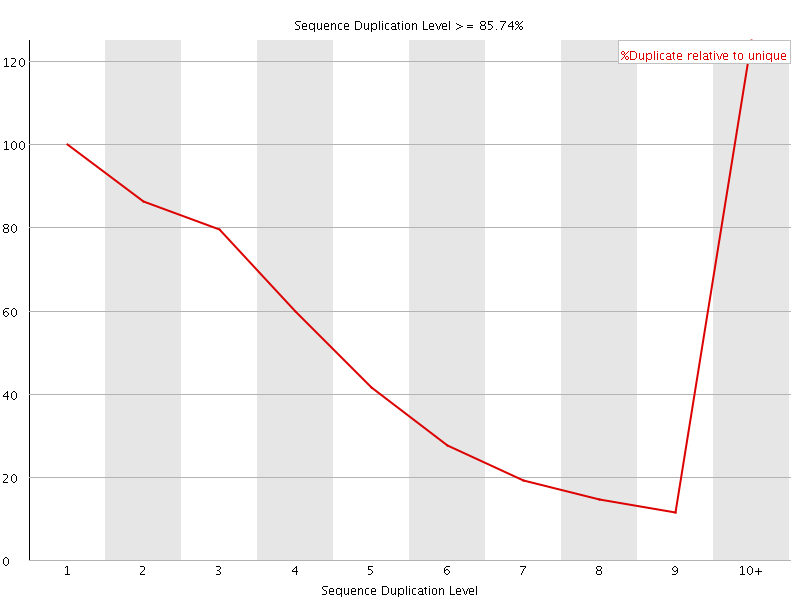
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
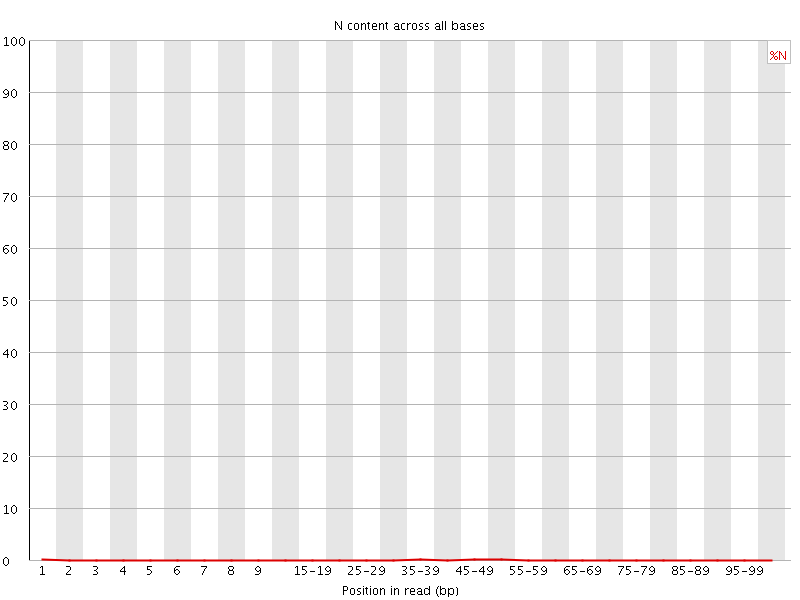
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
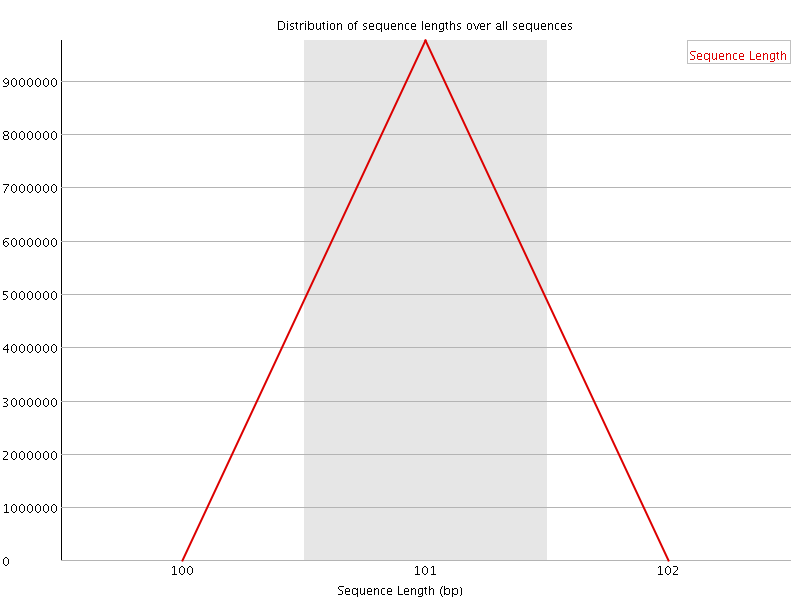
![[FAIL]](Icons/error.png) Sequence Duplication Levels
Sequence Duplication Levels
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCCTTATCTGTCAAAACCGCTAATGTCCGTTCTAAGACCGTCTGGAGA | 28726 | 0.2945471857393472 | No Hit |
| ACTGGATGCATCTGCAGGATATCGCGGCCGCTCGACGTAGAACTCAATCT | 28110 | 0.2882309194156182 | No Hit |
| ACTAGTATGGCCCGGGGGATCCTACGTTCCAAATGCAGCGAGCTCGTATA | 25055 | 0.2569059297743975 | No Hit |
| ACTGGATGCATCTGCAGGATATCGCGGCCGCCATCTGCCCTACGTTTGAG | 20727 | 0.21252800664274346 | No Hit |
| CTCCTGTTCACTTTTCTTCCCCAATTCGAGGCTCTACACGCAAATGGCTG | 17970 | 0.18425861337241764 | No Hit |
| ACTAGTATGGCCCGGGGGATCCTTATCTGTCAAAACCGCTAATGTCCGTT | 17860 | 0.18313070867175174 | No Hit |
| TCTGCAGGATATCGCGGCCGCCATCTGCCCTACGTTTGAGGGTTATAAGC | 15140 | 0.1552407015280135 | No Hit |
| GATCCTACGTTCCAAATGCAGCGAGCTCGTATAACCCTTTAAGAGTTGCT | 10435 | 0.10699714137680455 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
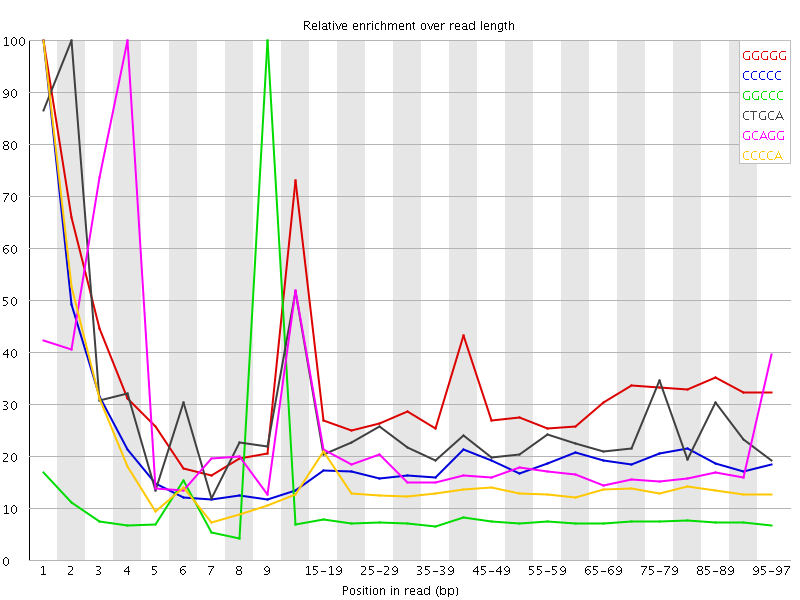
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GGGGG | 786285 | 2.949213 | 8.917768 | 1 |
| CCCCC | 720015 | 2.496202 | 12.932096 | 1 |
| GGCCC | 487970 | 1.7458481 | 20.485144 | 9 |
| CTGCA | 1155640 | 1.7367042 | 6.656066 | 2 |
| GCAGG | 720020 | 1.7094116 | 8.037061 | 4 |
| CCCCA | 743640 | 1.6840386 | 11.262022 | 1 |
| TGCAG | 1100590 | 1.6802211 | 5.58915 | 3 |
| CTGGG | 692490 | 1.644148 | 5.2276263 | 1 |
| TGGCC | 682215 | 1.5944504 | 14.393693 | 8 |
| GGCCG | 438150 | 1.5924792 | 5.898386 | 25-29 |
| CGGCC | 435700 | 1.5588379 | 5.8218493 | 25-29 |
| GCCCA | 667085 | 1.5346457 | 5.66173 | 1 |
| CCTGG | 643800 | 1.5046681 | 5.1210027 | 1 |
| CCCAG | 653200 | 1.5027027 | 6.488259 | 1 |
| CCTCC | 662120 | 1.4995165 | 5.7649336 | 1 |
| CCCAA | 1008690 | 1.4921006 | 5.272489 | 2 |
| CTGGA | 960030 | 1.4656345 | 11.507526 | 2 |
| GCGGC | 401100 | 1.4578191 | 5.3062134 | 20-24 |
| CCCTG | 632960 | 1.4562248 | 6.3076825 | 1 |
| CGGGG | 388690 | 1.4351324 | 6.157653 | 1 |
| GCCCC | 405830 | 1.4292887 | 7.7938137 | 1 |
| CCCCT | 626735 | 1.4193794 | 9.380249 | 1 |
| ACTGG | 918435 | 1.4021333 | 10.277606 | 1 |
| GCCGC | 388270 | 1.3891438 | 5.632214 | 25-29 |
| ATGGC | 905155 | 1.3818593 | 9.520418 | 7 |
| TGCAT | 1392435 | 1.3669562 | 7.371355 | 7 |
| CCAGT | 902845 | 1.3568022 | 5.646839 | 3 |
| ATGCA | 1367165 | 1.3420706 | 7.991444 | 6 |
| CTCCA | 903490 | 1.3365618 | 5.2596226 | 1 |
| CTCCC | 582610 | 1.3194486 | 8.570981 | 1 |
| GCCCT | 571050 | 1.313791 | 5.02537 | 1 |
| CGCGG | 341170 | 1.2400004 | 5.1124916 | 20-24 |
| CCTGT | 817695 | 1.2289094 | 7.442602 | 3 |
| CTCCT | 817420 | 1.209306 | 8.639414 | 1 |
| CCCGG | 334660 | 1.197339 | 5.596787 | 1 |
| TCCTG | 788390 | 1.1848673 | 6.6535397 | 2 |
| GCATC | 786010 | 1.1812215 | 9.9077015 | 8 |
| GATGC | 768690 | 1.1735244 | 10.5770235 | 5 |
| CATCT | 1179440 | 1.1397718 | 6.8998585 | 9 |
| TCCTT | 1167910 | 1.1286954 | 5.0365806 | 3 |
| CCCCG | 320235 | 1.1278323 | 7.533905 | 1 |
| GGATG | 704250 | 1.0922079 | 11.083866 | 4 |
| CCTCG | 430140 | 0.98960525 | 5.0410075 | 1 |
| TGGAT | 965685 | 0.9630587 | 7.851456 | 3 |
| CTCGG | 405060 | 0.9466929 | 5.2514954 | 1 |
| GATCC | 622585 | 0.93562526 | 8.744625 | 1 |
| CCCGC | 242980 | 0.8557488 | 5.0186133 | 1 |
| ATCCT | 841680 | 0.8133717 | 5.080686 | 2 |
| TATGG | 795110 | 0.7929476 | 5.7915664 | 6 |
| GTATG | 776675 | 0.7745627 | 6.00329 | 5 |
| CTAGT | 538795 | 0.52893615 | 5.680487 | 2 |
| ACTAG | 517790 | 0.50828594 | 5.5147424 | 1 |