![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 3291_5903_10009_H94MGADXX_V_CF26_TTAGGC_R2.fastq |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 19319381 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 36 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
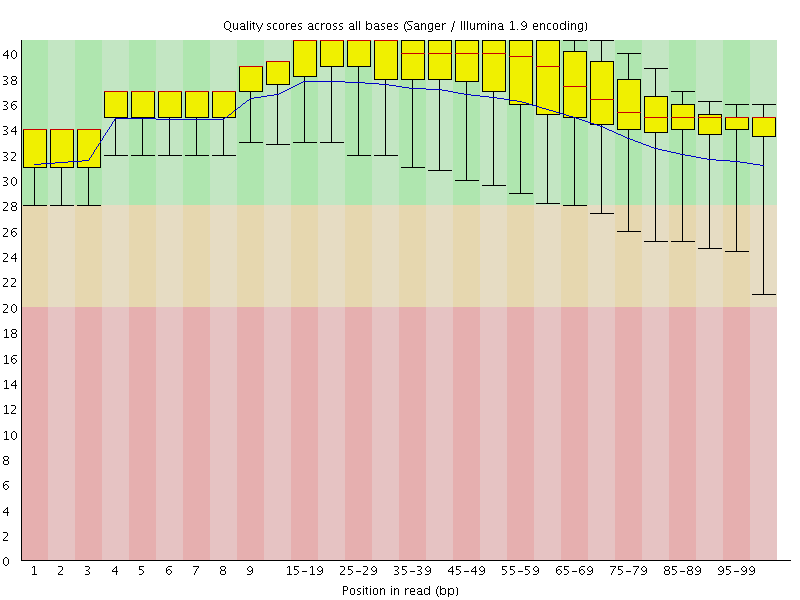
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
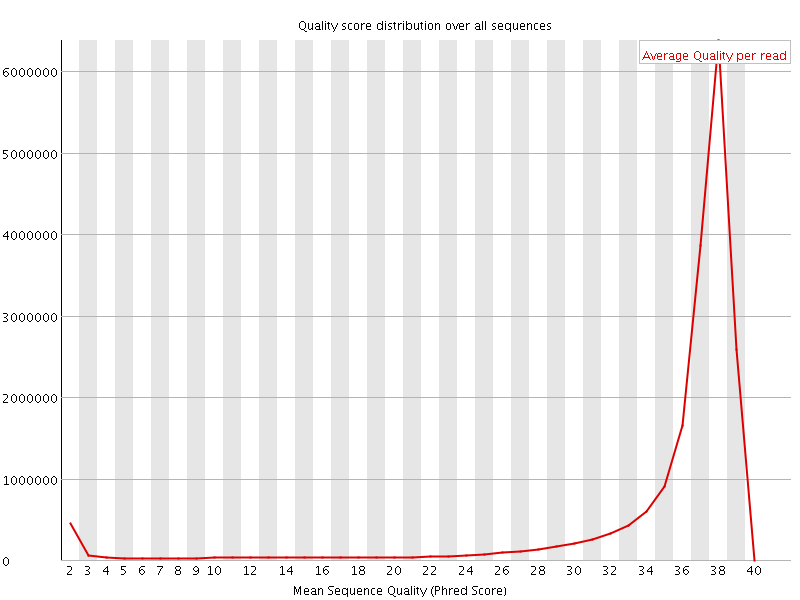
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
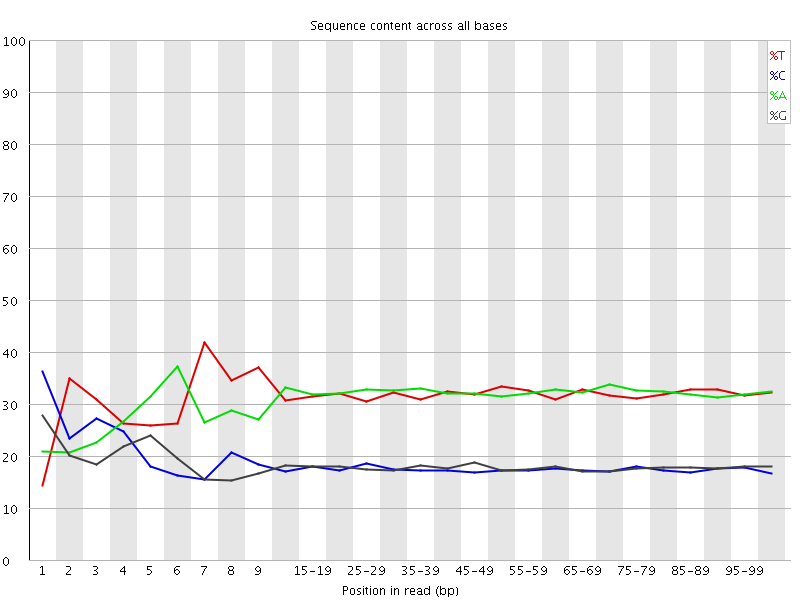
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
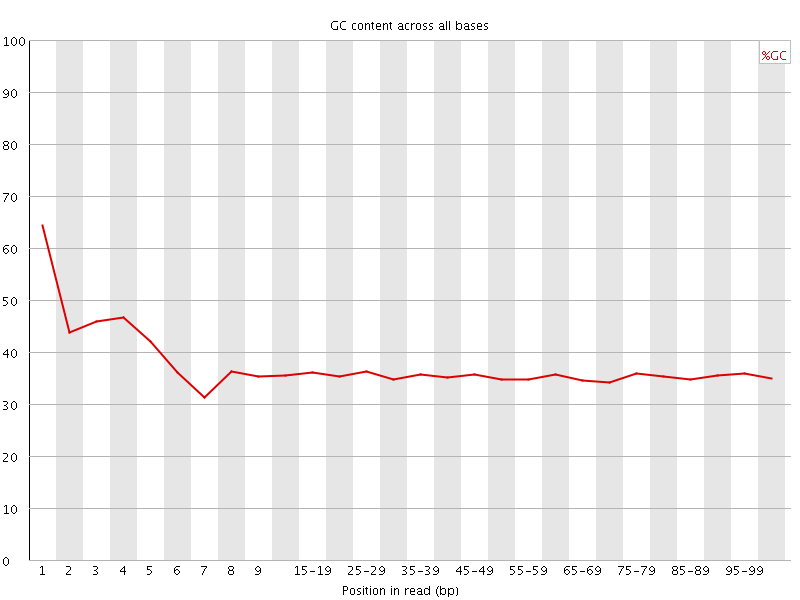
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
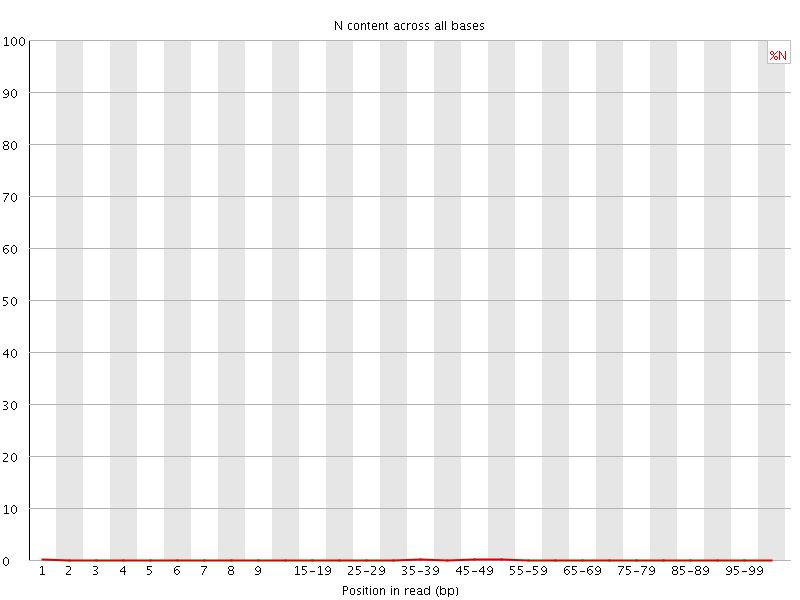
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
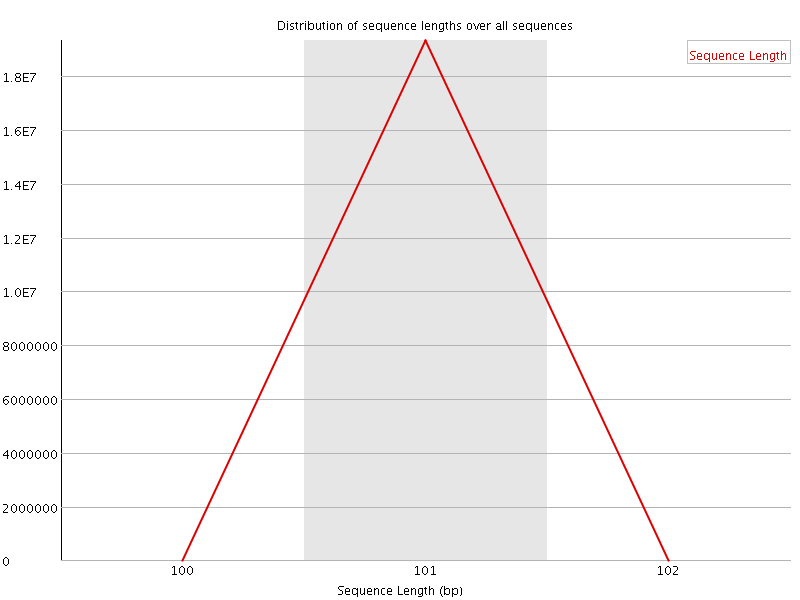
![[FAIL]](Icons/error.png) Sequence Duplication Levels
Sequence Duplication Levels
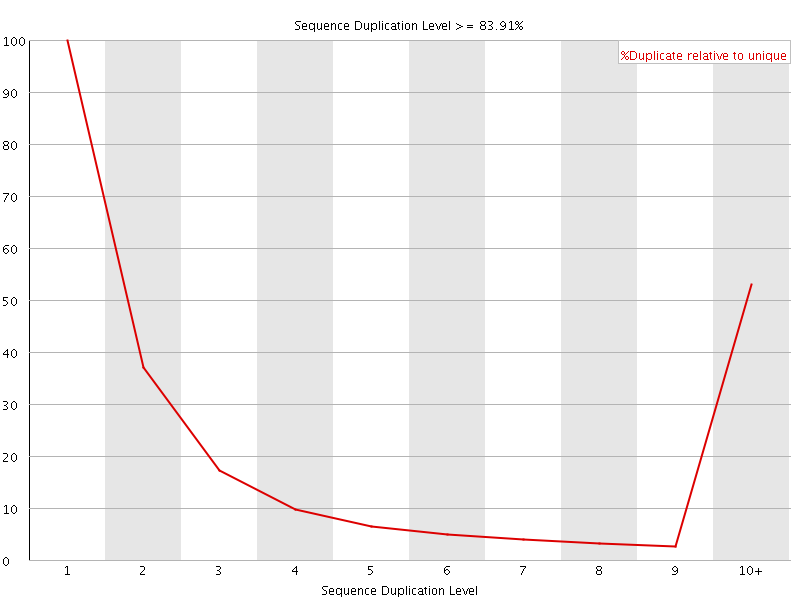
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GCCCCAACCAAAGTTATCTTATTAACTAACTAAATATAAATTATTAAAGC | 144086 | 0.745810644761341 | No Hit |
| CTTAGAAACTGCTGCATCTCTAGGAAATCTCGATCCAACATCGAGGTCGC | 110876 | 0.5739107272639843 | No Hit |
| GTCCTTTCGTACTAAGAAAAGTATAACTTTCGTGGATAGAAACTGACCTG | 107226 | 0.5550177824020345 | No Hit |
| CCCCAACCAAAGTTATCTTATTAACTAACTAAATATAAATTATTAAAGCT | 106475 | 0.5511304942948224 | No Hit |
| GCTGCATCTCTAGGAAATCTCGATCCAACATCGAGGTCGCAAACTTTTTT | 52790 | 0.2732489203458434 | No Hit |
| CTCTCCAAAAAAATTACGCTGTTATCCCTACGGTAACTTATTCCTTTGCT | 50188 | 0.259780579926448 | No Hit |
| CTTTGCTTAAAAGAAAAGCTTTAACGCTTTTATGCGAGTTAGATGCTATT | 47713 | 0.24696961046526286 | No Hit |
| CCCTACGGTAACTTATTCCTTTGCTCGCTATTTAGCGGATCACTTAATTT | 41325 | 0.2139043688822121 | No Hit |
| CTGCTGCATCTCTAGGAAATCTCGATCCAACATCGAGGTCGCAAACTTTT | 40795 | 0.21116100976527147 | No Hit |
| CCCCAGTCTTGCCATTCATACCAGCCTCTAATTAAGAGGCAAATGATTAT | 40277 | 0.20847976443965777 | No Hit |
| CTCCGCGATTGCCCCAACCAAAGTTATCTTATTAACTAACTAAATATAAA | 39872 | 0.20638342398237294 | No Hit |
| CCTTTGCTTAAAAGAAAAGCTTTAACGCTTTTATGCGAGTTAGATGCTAT | 39658 | 0.2052757280370422 | No Hit |
| GCCGAGTTCCTTTTTCTTATTTTATTAAAAAATTAATTACCTGAGTTGGA | 35079 | 0.1815741404965304 | No Hit |
| CGAGAAGACCCTATCGAGCTTTAGTAAAATTGTTAGTAGCATGTAAATAT | 33784 | 0.17487102718249617 | No Hit |
| CTGCATCTCTAGGAAATCTCGATCCAACATCGAGGTCGCAAACTTTTTTA | 33701 | 0.1744414067924847 | No Hit |
| CCTCGATGTTGGATCGAGATTTCCTAGAGATGCAGCAGTTTCTAAGGGTT | 33497 | 0.17338547233992643 | No Hit |
| CCCCAGTCTTGCCATTCATACCAGCCTCTATTTAAGAGGCAAATGATTAT | 33351 | 0.17262975454544843 | No Hit |
| GCTACCTTTGCACGGTCAAAATACCGCGGCCGTTTAACTACGTCACTGGG | 32833 | 0.16994850921983473 | No Hit |
| CAAAAACATCGCCCCTTGCAAAATAATTAACGAATAAGGGGTCCTGCCTG | 32242 | 0.16688940499698204 | No Hit |
| GGGCGATGTTTTTGGTAAACAGGCGAAGTCAATATTTGCCGAGTTCCTTT | 31822 | 0.1647154223005385 | No Hit |
| CTCTGGTCCTTTCGTACTAAGAAAAGTATAACTTTCGTGGATAGAAACTG | 31604 | 0.16358702175809878 | No Hit |
| GTCGCAAACTTTTTTATCAATATGAACTCTCCAAAAAAATTACGCTGTTA | 30272 | 0.15669239092080642 | No Hit |
| CCCTTAGAAACTGCTGCATCTCTAGGAAATCTCGATCCAACATCGAGGTC | 30037 | 0.1554759958406535 | No Hit |
| ACTGGATGCATCTGCAGGATATCGCGGCCGCTCGACGTAGAACTCAATCT | 29748 | 0.15398008869952923 | No Hit |
| GCTAAATAGCGAGCAAAGGAATAAGTTACCGTAGGGATAACAGCGTAATT | 27557 | 0.14263914563308214 | No Hit |
| ACTAGTATGGCCCGGGGGATCCTACGTTCCAAATGCAGCGAGCTCGTATA | 26141 | 0.13530971825650107 | No Hit |
| GCCCCGACCAAAGTTATCTTATTAACTAACTAAATATAAATTATTAAAGC | 25587 | 0.1324421315569065 | No Hit |
| CCCAACCAAAGTTATCTTATTAACTAACTAAATATAAATTATTAAAGCTT | 24892 | 0.1288447078092202 | No Hit |
| GAATAAGTTACCGTAGGGATAACAGCGTAATTTTTTTGGAGAGTTCATAT | 24293 | 0.1257441943921495 | No Hit |
| AGCTCATCAAGTTTCGGGACACAAAACACTTTTAGTAAAAGCTTGCTCAA | 23874 | 0.12357538784498322 | No Hit |
| ATTTTTGTGAAGAAGCAGAAATAAATTCGTAGGACGAGAAGACCCTATCG | 23472 | 0.12149457583553014 | No Hit |
| GCCGTTTAACTACGTCACTGGGCAGGCAGGACCCCTTATTCGTTAATTAT | 23254 | 0.1203661752930904 | No Hit |
| GCTACTAACAATTTTACTAAAGCTCGATAGGGTCTTCTCGTCCTACGAAT | 22783 | 0.11792820898350728 | No Hit |
| ACCAGCTATAACTTGATGCGATTAGCATTTCACATCTAATGTATCCTCAA | 22421 | 0.11605444294514404 | No Hit |
| CCCCGACCAAAGTTATCTTATTAACTAACTAAATATAAATTATTAAAGCT | 22248 | 0.11515896912018041 | No Hit |
| CACCATTAACTAAAATTAATTTCTTAGAAAACCAGCTATAACTTGATGCG | 21647 | 0.11204810340455523 | No Hit |
| AGATAACTTTGGTTGGGGCAATCGCGGAGAATATAAATCCTCCGCTAAAA | 21627 | 0.11194458041901031 | No Hit |
| AGAAGACCCTATCGAGCTTTAGTAAAATTGTTAGTAGCATGTAAATATAA | 20964 | 0.10851279344819588 | No Hit |
| CTGGGGTTAAGCTGTCTCTTTCTTAATACTTGAATTTTATATTTTTGTGA | 20608 | 0.10667008430549613 | No Hit |
| CTTTCGTACTAAGAAAAGTATAACTTTCGTGGATAGAAACTGACCTGGCT | 20137 | 0.10423211799591302 | No Hit |
| GGGCAGGCAGGACCCCTTATTCGTTAATTATTTTGCAAGGGGCGATGTTT | 20055 | 0.10380767375517881 | No Hit |
| GCTCGCTATTTAGCGGATCACTTAATTTAAAAATAGTAATTTTTAATACT | 19810 | 0.1025395171822534 | No Hit |
| CTGGTCCTTTCGTACTAAGAAAAGTATAACTTTCGTGGATAGAAACTGAC | 19694 | 0.10193908386609281 | No Hit |
| TGCCCCAACCAAAGTTATCTTATTAACTAACTAAATATAAATTATTAAAG | 19585 | 0.10137488359487294 | No Hit |
| AAAGTATTAAAAATTACTATTTTTAAATTAAGTGATCCGCTAAATAGCGA | 19503 | 0.10095043935413872 | No Hit |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
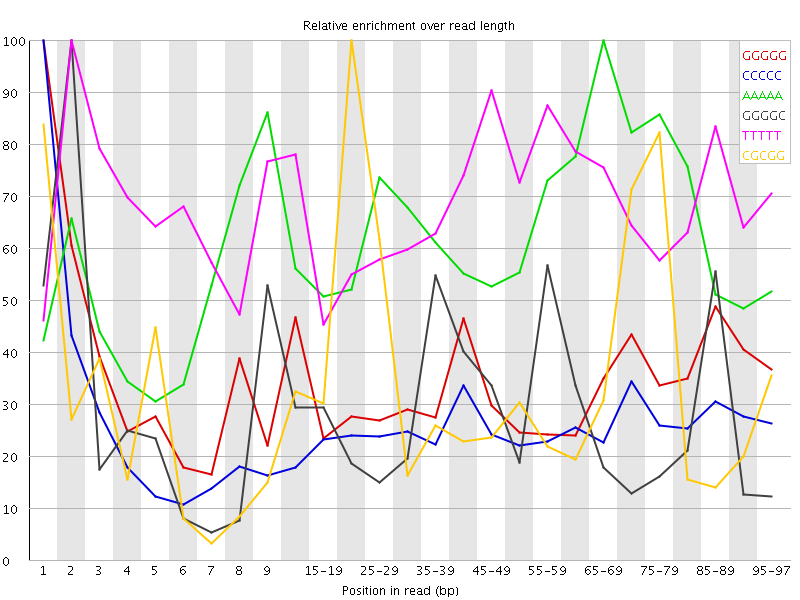
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GGGGG | 1339350 | 3.6911366 | 10.8413 | 1 |
| CCCCC | 1151080 | 3.2541585 | 12.599269 | 1 |
| AAAAA | 19823395 | 3.148251 | 4.906727 | 65-69 |
| GGGGC | 1099465 | 3.0455158 | 10.662783 | 2 |
| TTTTT | 18446580 | 3.0002127 | 4.3543878 | 2 |
| CGCGG | 990695 | 2.758245 | 7.741347 | 20-24 |
| GCCCC | 973485 | 2.738098 | 54.117176 | 1 |
| GCAGG | 1599015 | 2.502978 | 8.669151 | 2 |
| CCCCT | 1565725 | 2.5005138 | 8.051741 | 1 |
| CCAAC | 2673225 | 2.388867 | 17.259506 | 4 |
| CTGGG | 1517240 | 2.3863149 | 11.913057 | 1 |
| TGGGG | 1505915 | 2.3564622 | 8.316163 | 8 |
| CCCCA | 1478805 | 2.3504753 | 35.9445 | 1 |
| GCCGG | 837035 | 2.3304322 | 9.355244 | 55-59 |
| GACCC | 1462300 | 2.3124259 | 6.7409425 | 7 |
| CCGCG | 818730 | 2.2911158 | 12.782531 | 3 |
| GGCAG | 1458050 | 2.282322 | 8.43903 | 1 |
| GTTGG | 2522500 | 2.241224 | 5.8112087 | 8 |
| TCCGC | 1375755 | 2.185956 | 9.438307 | 2 |
| GGCCG | 782900 | 2.1797123 | 7.607628 | 75-79 |
| TTCCT | 4254760 | 2.168453 | 5.8272495 | 65-69 |
| GGGCA | 1338755 | 2.0955865 | 6.2078733 | 1 |
| GCTGC | 1311825 | 2.0737808 | 10.861877 | 1 |
| CGGCC | 739645 | 2.0698059 | 7.8504066 | 75-79 |
| GGGGT | 1314385 | 2.0567555 | 5.883766 | 3 |
| CCTGC | 1287755 | 2.0461316 | 6.9694414 | 1 |
| GGGCG | 738360 | 2.0452557 | 11.457464 | 1 |
| CAGGC | 1294840 | 2.037202 | 5.8875804 | 20-24 |
| CCCTA | 2223180 | 1.9961816 | 8.296202 | 1 |
| CCTTT | 3904560 | 1.9899724 | 9.327375 | 3 |
| CCCTT | 2196240 | 1.9814091 | 5.849352 | 1 |
| GGGGA | 1252585 | 1.9507351 | 5.7347717 | 1 |
| CTTCC | 2160270 | 1.9489576 | 5.255876 | 1 |
| GGCGA | 1239865 | 1.9407917 | 7.3989415 | 2 |
| TGGGC | 1219065 | 1.9173452 | 6.8860865 | 95-97 |
| CCTCC | 1197070 | 1.9117597 | 6.166152 | 1 |
| CCGGC | 668835 | 1.8716531 | 5.383936 | 1 |
| TCGAG | 2081650 | 1.850147 | 5.2568865 | 30-34 |
| CCCAG | 1166250 | 1.8442639 | 16.149927 | 2 |
| CGCCG | 648950 | 1.8160069 | 9.783821 | 55-59 |
| CCAAA | 3589310 | 1.8033459 | 11.386059 | 8 |
| GGTCG | 1138590 | 1.7907742 | 6.9733996 | 45-49 |
| AGCTT | 3491950 | 1.7622218 | 5.0510764 | 45-49 |
| CGAGG | 1124320 | 1.7599261 | 6.975988 | 30-34 |
| CAACC | 1945725 | 1.738753 | 16.324749 | 5 |
| CTTTC | 3396215 | 1.7308927 | 7.466169 | 4 |
| TCTCC | 1882345 | 1.6982186 | 5.4683867 | 2 |
| AAGCT | 3365450 | 1.6903114 | 5.3934536 | 45-49 |
| GCCTG | 1065535 | 1.6844364 | 5.615513 | 1 |
| CGCTG | 1050540 | 1.6607318 | 5.904078 | 1 |
| CTGGC | 1048490 | 1.6574912 | 6.0565724 | 45-49 |
| TTGCC | 1836315 | 1.6482692 | 12.79931 | 9 |
| GCCCA | 1031745 | 1.6315624 | 5.1352544 | 1 |
| CTCTC | 1804295 | 1.6278032 | 6.9644737 | 1 |
| CCTGG | 1021435 | 1.6147215 | 5.5870285 | 45-49 |
| CGCCC | 572800 | 1.6111008 | 7.2316456 | 1 |
| CAAAG | 3186020 | 1.5925868 | 11.152719 | 9 |
| CCTAC | 1758185 | 1.578665 | 5.2918 | 2 |
| ACCAA | 3126435 | 1.5707875 | 11.470088 | 7 |
| TCCTA | 3094935 | 1.5698478 | 5.4278626 | 65-69 |
| CCTCG | 987280 | 1.5687028 | 9.938493 | 1 |
| CTGCT | 1741450 | 1.5631187 | 11.318887 | 9 |
| TCCTT | 3056220 | 1.557613 | 8.831912 | 2 |
| CCAGC | 982990 | 1.5544634 | 5.979183 | 2 |
| AAAGT | 5489085 | 1.5500124 | 5.242407 | 9 |
| CCGGG | 552590 | 1.5384946 | 9.19794 | 1 |
| CTCCG | 966455 | 1.5356137 | 10.162965 | 1 |
| CCCAA | 1692165 | 1.5121648 | 15.827765 | 3 |
| CGGGG | 543950 | 1.5067405 | 10.088486 | 1 |
| CCTAG | 1683000 | 1.5034746 | 8.215201 | 65-69 |
| CTCCT | 1659610 | 1.497271 | 6.61014 | 1 |
| CTTTG | 2951545 | 1.4966178 | 5.6670256 | 1 |
| GCGGC | 536370 | 1.4933355 | 6.9723105 | 75-79 |
| CTCCA | 1632240 | 1.4655796 | 5.7182417 | 1 |
| CCCCG | 512975 | 1.4428326 | 14.434618 | 1 |
| CCCGG | 513710 | 1.4375545 | 6.2669935 | 1 |
| CTCGC | 900960 | 1.4315478 | 5.668119 | 1 |
| TCCTG | 1590050 | 1.4272226 | 5.6703243 | 2 |
| CTGCA | 1592215 | 1.4223735 | 7.825463 | 2 |
| CAGCC | 894030 | 1.4137852 | 5.1673617 | 20-24 |
| CTCCC | 882365 | 1.4091656 | 7.643144 | 1 |
| TCGCC | 875320 | 1.390808 | 6.7825274 | 9 |
| TCCCT | 1536140 | 1.3858786 | 5.8145847 | 95-97 |
| ATCCC | 1542565 | 1.3850608 | 6.120023 | 95-97 |
| CTTGC | 1541685 | 1.3838103 | 9.134736 | 8 |
| CGGTC | 873225 | 1.3804259 | 6.404762 | 60-64 |
| GTCCT | 1512995 | 1.3580583 | 14.66735 | 1 |
| GCTCG | 854600 | 1.3509827 | 5.161943 | 1 |
| CCGAG | 850100 | 1.3374821 | 6.9417768 | 2 |
| TGCTG | 1448370 | 1.2934422 | 5.8350024 | 2 |
| GAAAC | 2581180 | 1.2902473 | 6.601098 | 5 |
| CTCGG | 815230 | 1.2887452 | 8.849594 | 1 |
| CCGGT | 805040 | 1.2726367 | 5.7033396 | 55-59 |
| ACTGG | 1428815 | 1.2699146 | 5.3314567 | 1 |
| GCATC | 1418220 | 1.2669386 | 6.1732907 | 4 |
| GTCGC | 793710 | 1.2547257 | 6.8119917 | 45-49 |
| GGCCC | 448200 | 1.2542328 | 12.007685 | 9 |
| AACCA | 2472030 | 1.2420006 | 10.355564 | 6 |
| CCCTG | 765355 | 1.216083 | 5.5724673 | 1 |
| ACTGC | 1353760 | 1.2093545 | 11.054088 | 8 |
| TGGCC | 763315 | 1.2066761 | 7.8582296 | 8 |
| GTGGG | 769665 | 1.2043751 | 5.2455764 | 1 |
| GCCGA | 752935 | 1.18461 | 6.9484124 | 1 |
| CGATG | 1329290 | 1.181458 | 7.0236325 | 4 |
| ATGGC | 1317875 | 1.1713125 | 5.223432 | 7 |
| ACGCC | 731705 | 1.1570905 | 5.6169586 | 55-59 |
| GATGC | 1298420 | 1.154021 | 5.479631 | 5 |
| GTCTT | 2272100 | 1.1520969 | 5.9635644 | 6 |
| GCTAC | 1282520 | 1.1457137 | 7.43333 | 95-97 |
| CGTGG | 720455 | 1.1331315 | 5.468548 | 30-34 |
| GGTCC | 707775 | 1.1188765 | 6.170765 | 5 |
| TTCGT | 2196580 | 1.1138034 | 7.2659736 | 6 |
| GATGT | 2185305 | 1.0972135 | 5.1191754 | 5 |
| CATGC | 1217965 | 1.0880448 | 7.8110113 | 90-94 |
| AACTG | 2141360 | 1.075507 | 6.401281 | 7 |
| CTACT | 2106370 | 1.0684166 | 6.608842 | 95-97 |
| CGGGC | 372375 | 1.0367485 | 6.682529 | 1 |
| TCGGC | 655140 | 1.0356692 | 5.076784 | 2 |
| GCCGT | 647105 | 1.0229671 | 6.181899 | 1 |
| CCAGT | 1141455 | 1.0196961 | 10.000706 | 3 |
| GCGAT | 1101300 | 0.97882307 | 6.522468 | 5 |
| GCCCT | 591925 | 0.9405177 | 5.5878944 | 1 |
| GTACT | 1820405 | 0.9186723 | 6.838634 | 9 |
| GCTCA | 1017680 | 0.9091242 | 5.0605035 | 2 |
| CCCGA | 570505 | 0.9021751 | 5.6357636 | 2 |
| TCTTG | 1749045 | 0.88687515 | 5.938412 | 7 |
| CTGGA | 948430 | 0.84295386 | 5.651307 | 2 |
| CGGGA | 523440 | 0.8193537 | 5.878824 | 1 |
| CTCTG | 880435 | 0.790275 | 5.0003815 | 1 |
| CCCGC | 280655 | 0.78939164 | 6.5079355 | 1 |
| GGATG | 872470 | 0.77149945 | 5.396371 | 4 |
| TCGTA | 1502780 | 0.7583819 | 7.253103 | 7 |
| TTTCG | 1464340 | 0.7425119 | 6.324889 | 5 |
| CTTAG | 1407560 | 0.7103289 | 6.779087 | 1 |
| CAGTC | 780910 | 0.6976104 | 8.37154 | 4 |
| CGCGA | 435655 | 0.6854261 | 6.878741 | 4 |
| CGTAC | 554175 | 0.4950612 | 10.405604 | 8 |