![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 3291_5903_10010_H94MGADXX_HK_CF2_TGACCA_R1.fastq |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 23172864 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
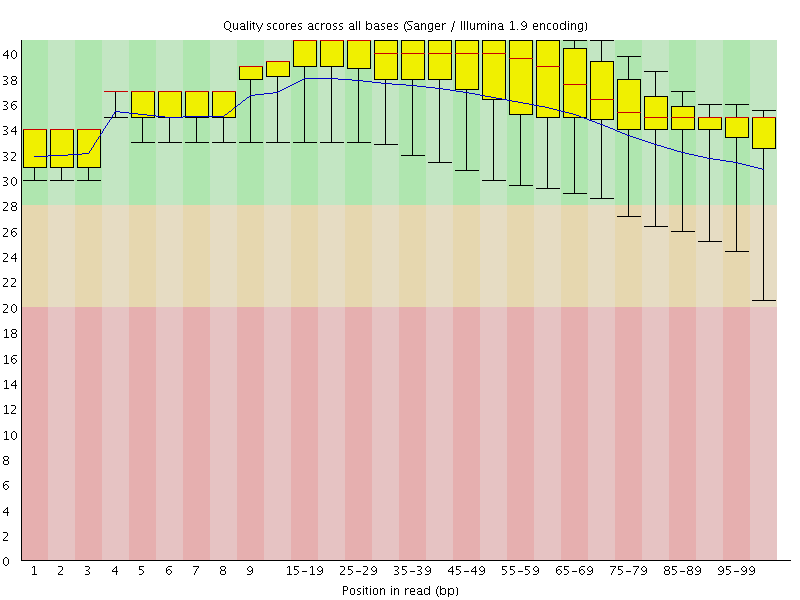
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
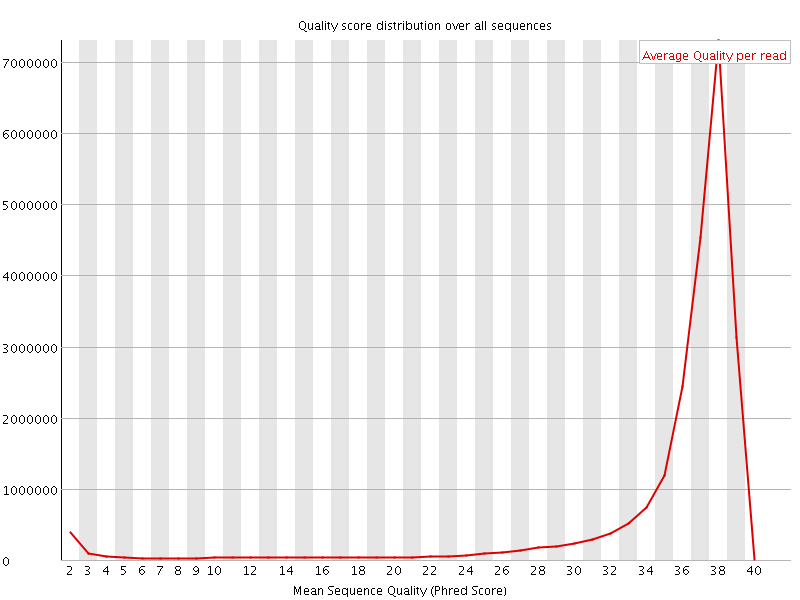
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
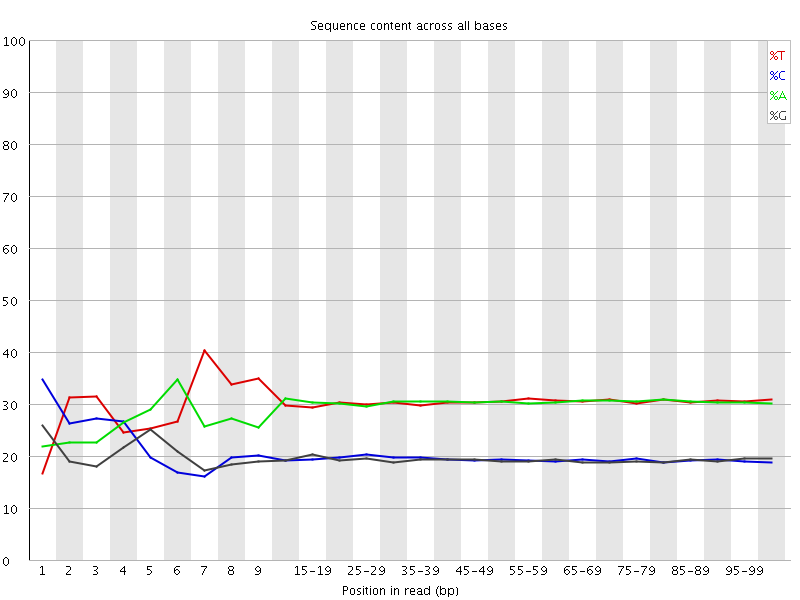
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
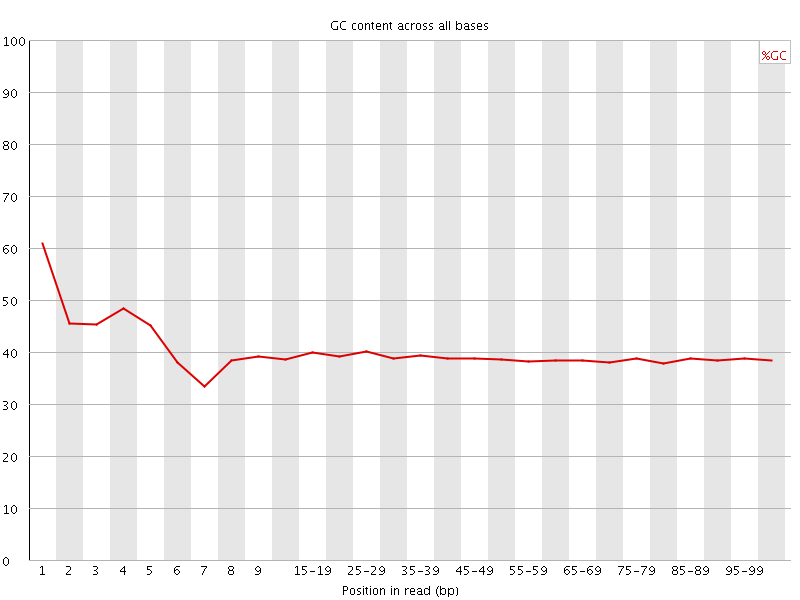
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
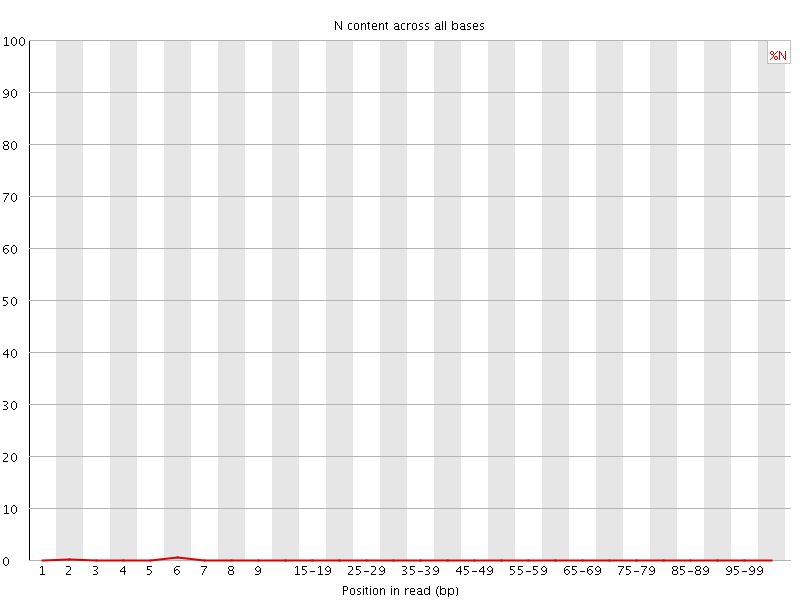
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
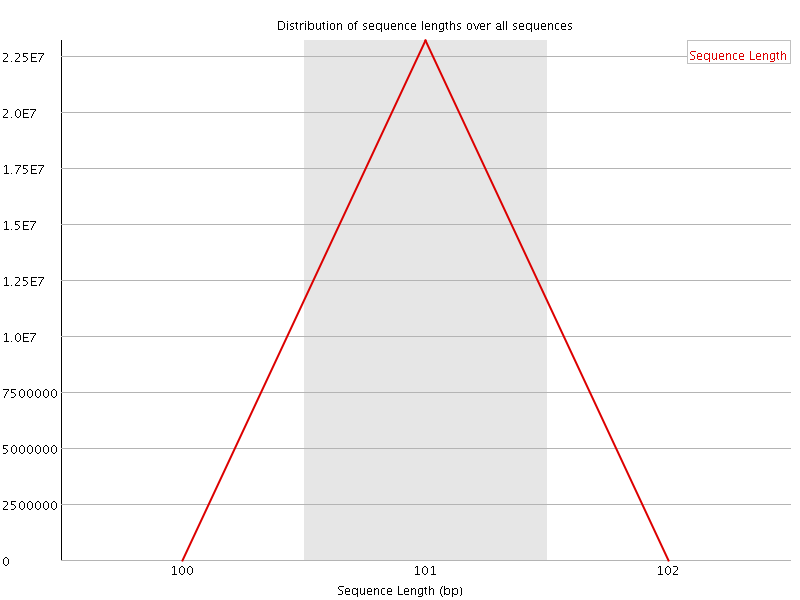
![[FAIL]](Icons/error.png) Sequence Duplication Levels
Sequence Duplication Levels
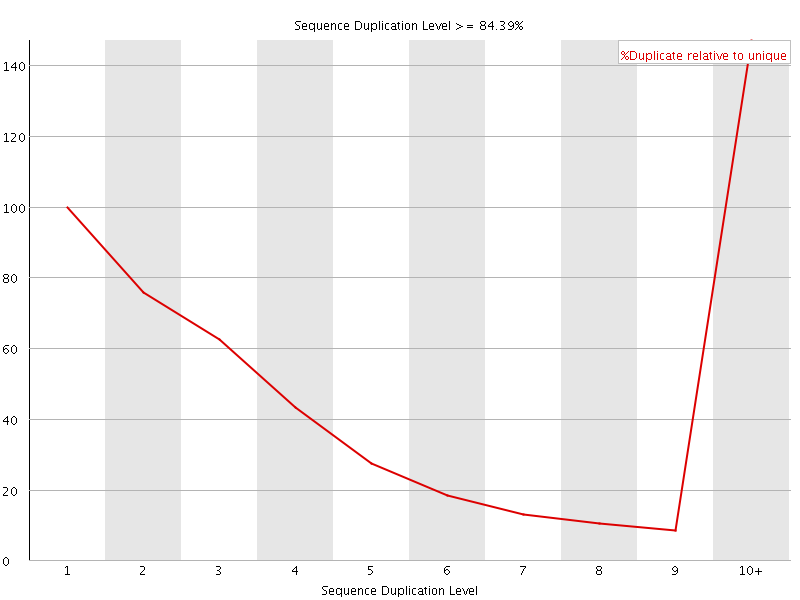
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| ACTAGTATGGCCCGGGGGATCCTACGTTCCAAATGCAGCGAGCTCGTATA | 155891 | 0.672730828610568 | No Hit |
| ACTGGATGCATCTGCAGGATATCGCGGCCGCTCGACGTAGAACTCAATCT | 139621 | 0.6025193950993715 | No Hit |
| GATCCTTATCTGTCAAAACCGCTAATGTCCGTTCTAAGACCGTCTGGAGA | 77254 | 0.3333813204962494 | No Hit |
| GATCCTACGTTCCAAATGCAGCGAGCTCGTATAACCCTTTAAGAGTTGCT | 74393 | 0.32103498298699723 | No Hit |
| ACTAGTATGGCCCGGGGGATCCTTATCTGTCAAAACCGCTAATGTCCGTT | 66005 | 0.2848374719672113 | No Hit |
| TCTGCAGGATATCGCGGCCGCCATCTGCCCTACGTTTGAGGGTTATAAGC | 62344 | 0.26903882057910494 | No Hit |
| ACTGGATGCATCTGCAGGATATCGCGGCCGCCATCTGCCCTACGTTTGAG | 61587 | 0.2657720685712392 | No Hit |
| TCTGCAGGATATCGCGGCCGCTCGACGTAGAACTCAATCTAAAACTTCGA | 47907 | 0.206737501242833 | No Hit |
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACTGACCAATCTCGTATGC | 32998 | 0.1423993167180371 | TruSeq Adapter, Index 4 (100% over 50bp) |
| CTGCAGGATATCGCGGCCGCCATCTGCCCTACGTTTGAGGGTTATAAGCT | 25885 | 0.11170393094267503 | No Hit |
| CTGCAGGATATCGCGGCCGCTCGACGTAGAACTCAATCTAAAACTTCGAT | 24597 | 0.10614570559771981 | No Hit |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
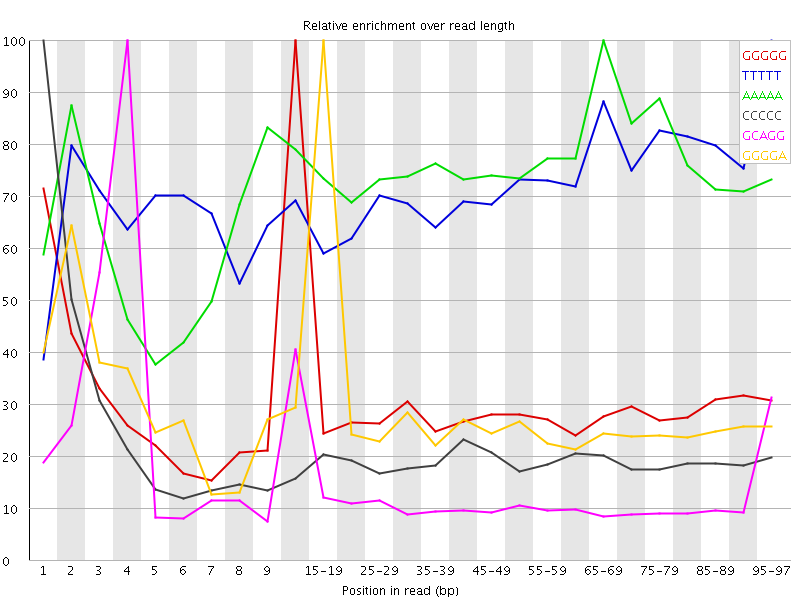
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GGGGG | 2242150 | 3.5354156 | 11.133234 | 10-14 |
| TTTTT | 19341225 | 3.2959123 | 4.5382895 | 95-97 |
| AAAAA | 18305505 | 3.2368717 | 4.285457 | 65-69 |
| CCCCC | 1671895 | 2.4199388 | 12.141609 | 1 |
| GCAGG | 1996035 | 1.9973836 | 14.556551 | 4 |
| GGGGA | 1845765 | 1.8789093 | 6.4011583 | 15-19 |
| TGCAG | 2924735 | 1.8755256 | 9.889176 | 3 |
| GGCCG | 1217070 | 1.8544674 | 8.533432 | 25-29 |
| CGGCC | 1228200 | 1.8396561 | 8.379994 | 25-29 |
| CTGCA | 2891100 | 1.8224831 | 10.962679 | 2 |
| GGCCC | 1194775 | 1.7895906 | 38.22523 | 9 |
| GCGGC | 1146215 | 1.7465045 | 8.01582 | 20-24 |
| CCCCA | 1782990 | 1.6948606 | 11.052858 | 1 |
| GCCGC | 1129060 | 1.6911596 | 8.128736 | 25-29 |
| TGGCC | 1664190 | 1.6249865 | 25.960999 | 8 |
| CGGGG | 1024295 | 1.5876865 | 9.173176 | 10-14 |
| CCTCC | 1677275 | 1.5826283 | 5.65527 | 1 |
| CAGGA | 2424755 | 1.5664437 | 9.276418 | 5 |
| GCCCA | 1618815 | 1.565375 | 5.226167 | 1 |
| TCTGC | 2489770 | 1.5579345 | 9.073196 | 1 |
| CGCGG | 1020220 | 1.5545242 | 7.668636 | 20-24 |
| CCTGG | 1581665 | 1.5444055 | 5.0892334 | 1 |
| CCCAG | 1577700 | 1.5256172 | 6.661319 | 1 |
| CTGGA | 2317110 | 1.4858779 | 16.544687 | 2 |
| CCGGG | 950250 | 1.4479098 | 8.408848 | 10-14 |
| CTCCA | 2329345 | 1.4434379 | 5.3231626 | 1 |
| CCCGG | 955720 | 1.4315227 | 8.397833 | 10-14 |
| CCCTG | 1455890 | 1.3974597 | 6.4073467 | 1 |
| ACTGG | 2172555 | 1.3931801 | 15.451548 | 1 |
| GCCCC | 940700 | 1.3851048 | 7.841817 | 1 |
| ATGCA | 3328605 | 1.3780098 | 11.346413 | 6 |
| ATGGC | 2145735 | 1.3759813 | 17.166014 | 7 |
| CCCCT | 1441475 | 1.3601342 | 8.317033 | 1 |
| CCCAC | 1405140 | 1.335687 | 5.272423 | 1 |
| TGCAT | 3245170 | 1.3335739 | 10.596764 | 7 |
| CTCCC | 1396125 | 1.3173431 | 6.9730744 | 1 |
| GCCCG | 868720 | 1.3012099 | 8.227605 | 10-14 |
| CCAGT | 2062725 | 1.3002946 | 5.019667 | 3 |
| CTCCT | 2084020 | 1.2819047 | 6.480139 | 1 |
| TCCTT | 2924550 | 1.1727142 | 5.3146577 | 3 |
| CCTAC | 1874820 | 1.16178 | 5.8485928 | 4 |
| CATCT | 2867095 | 1.1582056 | 10.079071 | 9 |
| GATGC | 1791655 | 1.148923 | 15.727017 | 5 |
| GCATC | 1811860 | 1.1421549 | 14.963738 | 8 |
| GGATG | 1724455 | 1.1249274 | 16.172564 | 4 |
| GTTCC | 1760955 | 1.10189 | 5.316969 | 9 |
| CCCCG | 746005 | 1.0984322 | 7.329865 | 1 |
| GATCC | 1697640 | 1.0701532 | 11.810511 | 1 |
| AGGAT | 2315555 | 0.97517216 | 5.948079 | 6 |
| TGGAT | 2310615 | 0.96592486 | 11.128596 | 3 |
| ATCCT | 2329160 | 0.9408987 | 7.3463454 | 2 |
| TCGCG | 940280 | 0.9181296 | 5.1900835 | 20-24 |
| TATGG | 2058310 | 0.8604518 | 10.791726 | 6 |
| CTACG | 1342260 | 0.8461299 | 5.190079 | 5 |
| GGATA | 1980915 | 0.834242 | 5.8606973 | 7 |
| AGTAT | 2930000 | 0.7907459 | 7.214298 | 4 |
| GTATG | 1870730 | 0.78203624 | 10.970614 | 5 |
| CGTTC | 1231360 | 0.77050424 | 5.098425 | 8 |
| TAGTA | 2429005 | 0.6555378 | 6.936389 | 3 |
| CTAGT | 1409685 | 0.57929766 | 10.627083 | 2 |
| ACTAG | 1368745 | 0.56664705 | 10.599375 | 1 |