![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 3291_5903_10010_H94MGADXX_HK_CF2_TGACCA_R2.fastq |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 23172864 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
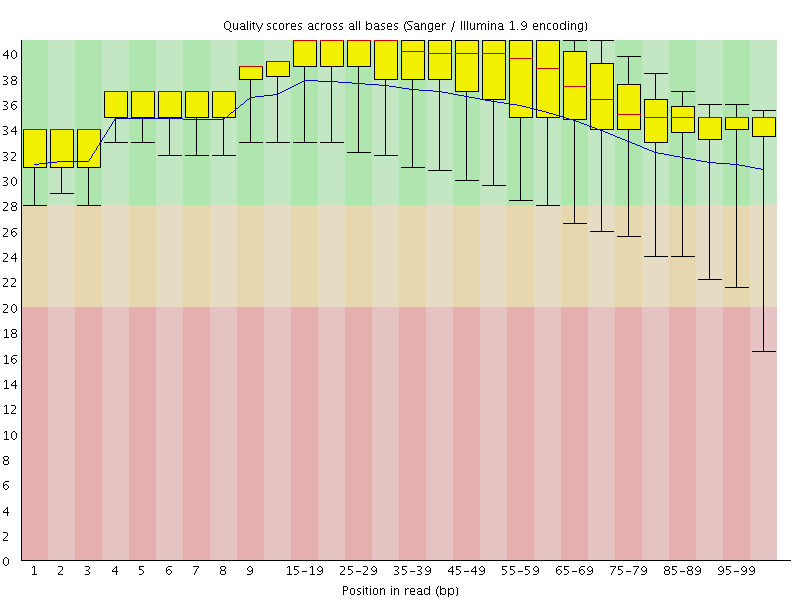
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
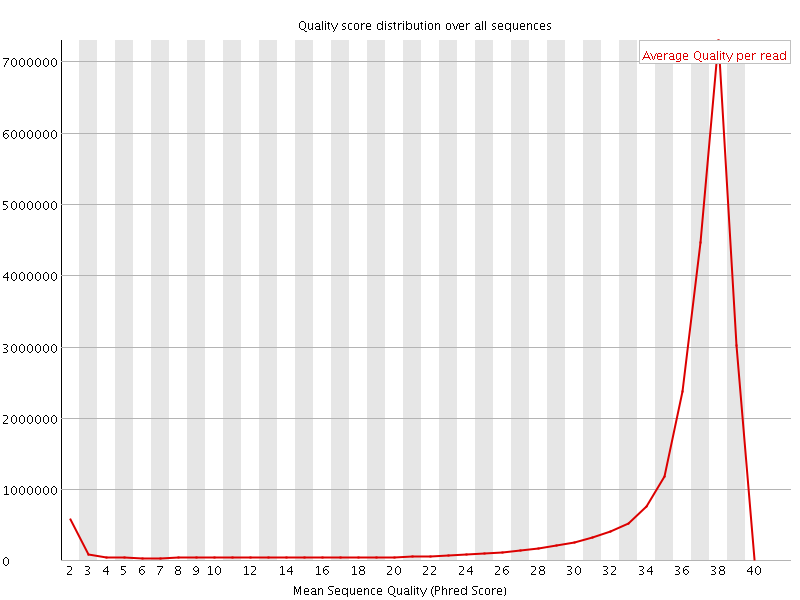
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
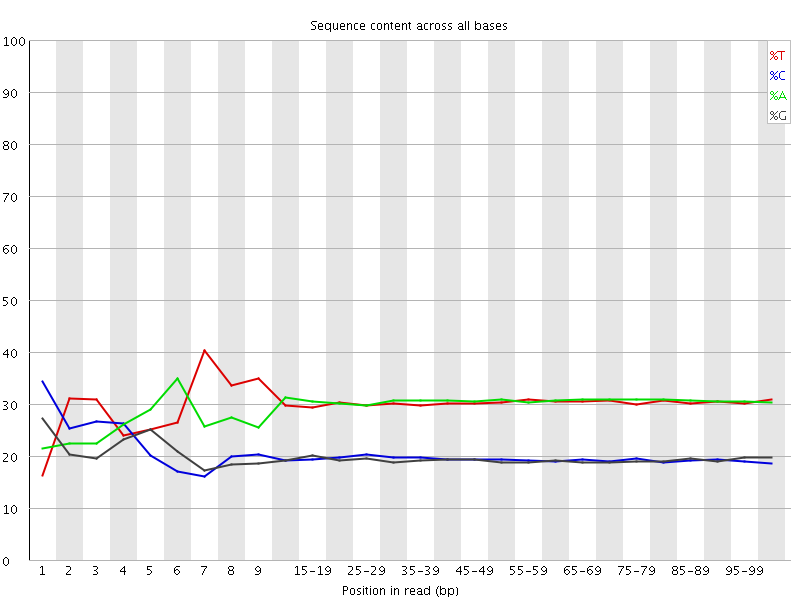
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
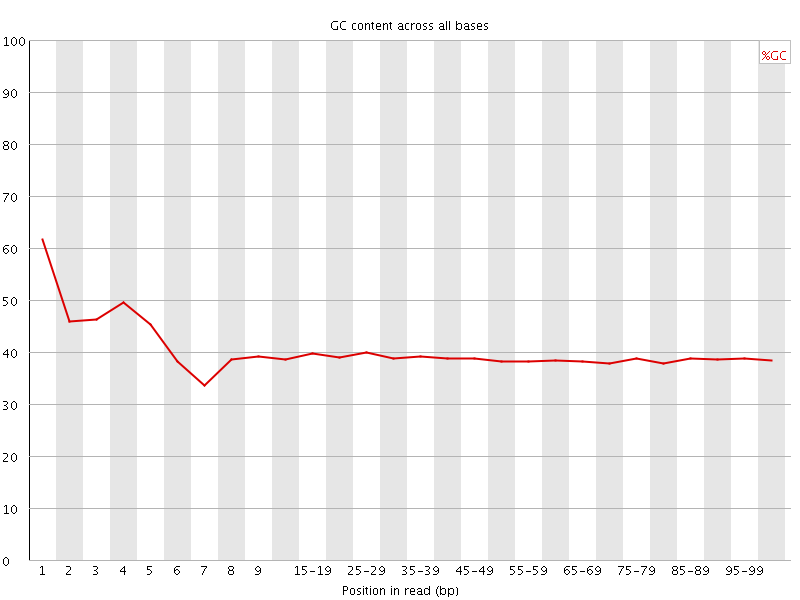
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
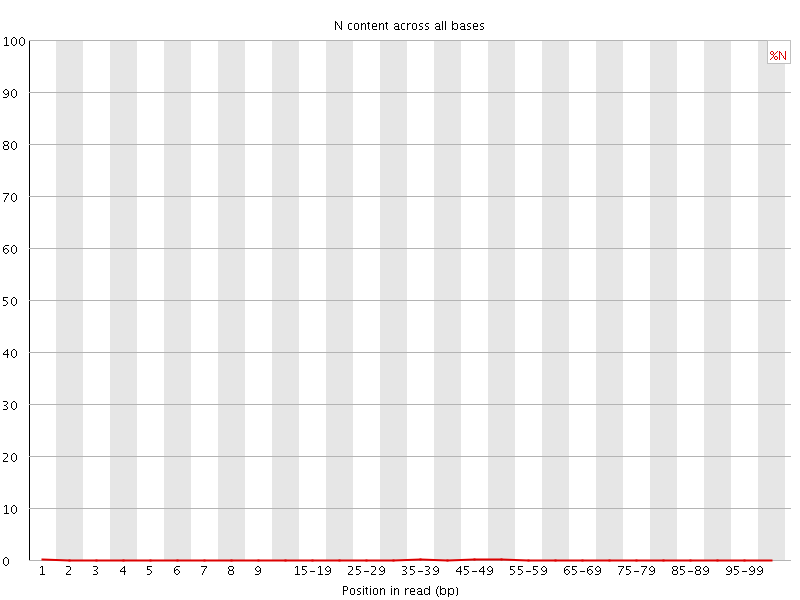
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
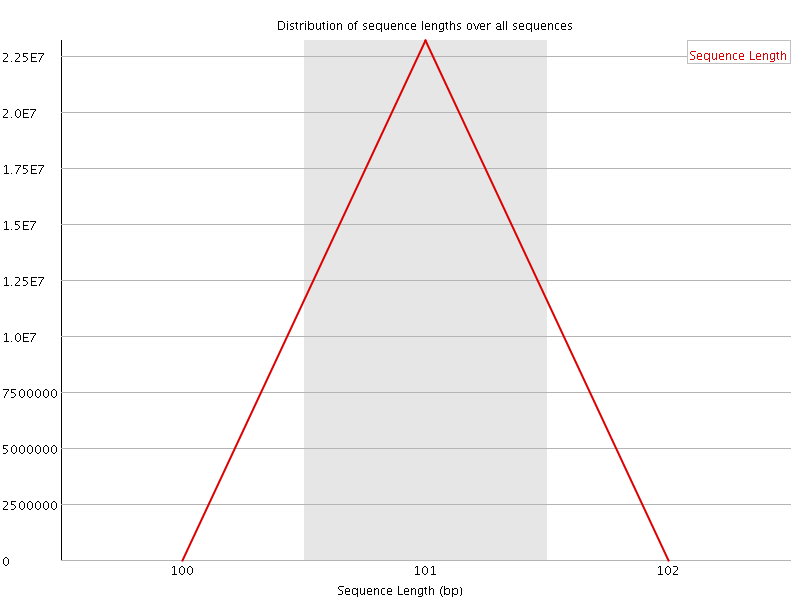
![[FAIL]](Icons/error.png) Sequence Duplication Levels
Sequence Duplication Levels
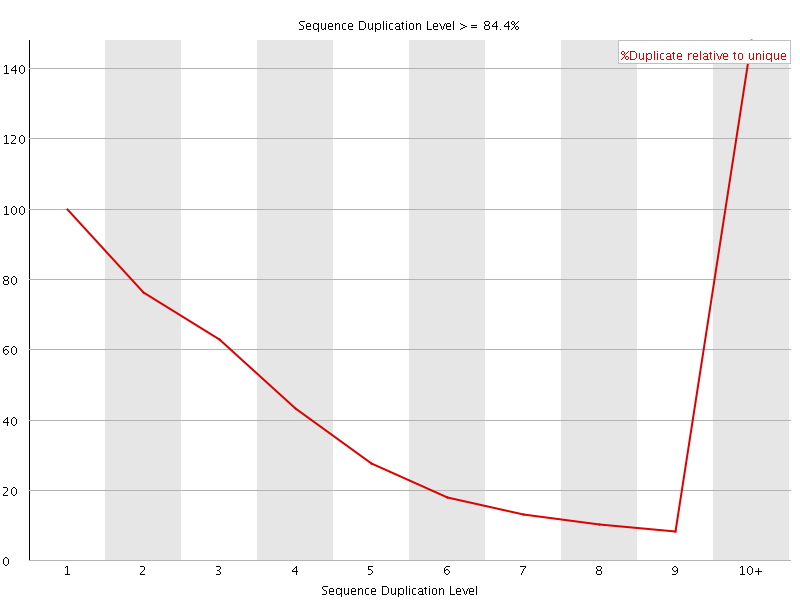
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| ACTGGATGCATCTGCAGGATATCGCGGCCGCTCGACGTAGAACTCAATCT | 156718 | 0.6762996580828334 | No Hit |
| ACTAGTATGGCCCGGGGGATCCTACGTTCCAAATGCAGCGAGCTCGTATA | 137922 | 0.5951875434991548 | No Hit |
| GATCCTTATCTGTCAAAACCGCTAATGTCCGTTCTAAGACCGTCTGGAGA | 94192 | 0.40647543609628917 | No Hit |
| GATCCTACGTTCCAAATGCAGCGAGCTCGTATAACCCTTTAAGAGTTGCT | 77393 | 0.33398116003270034 | No Hit |
| ACTGGATGCATCTGCAGGATATCGCGGCCGCCATCTGCCCTACGTTTGAG | 66861 | 0.2885314478175853 | No Hit |
| ACTAGTATGGCCCGGGGGATCCTTATCTGTCAAAACCGCTAATGTCCGTT | 59600 | 0.25719738397463515 | No Hit |
| TCTGCAGGATATCGCGGCCGCCATCTGCCCTACGTTTGAGGGTTATAAGC | 47641 | 0.20558960687811398 | No Hit |
| TCTGCAGGATATCGCGGCCGCTCGACGTAGAACTCAATCTAAAACTTCGA | 43031 | 0.1856956481512169 | No Hit |
| CTGCAGGATATCGCGGCCGCTCGACGTAGAACTCAATCTAAAACTTCGAT | 25938 | 0.1119326467371491 | No Hit |
| CTGCAGGATATCGCGGCCGCCATCTGCCCTACGTTTGAGGGTTATAAGCT | 23441 | 0.10115711204277554 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
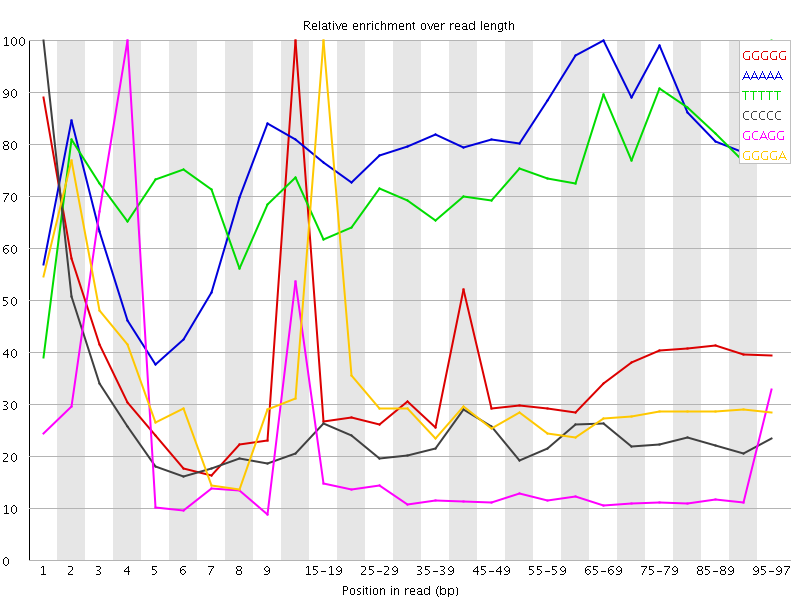
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GGGGG | 2388195 | 3.7501304 | 9.967601 | 10-14 |
| AAAAA | 20622330 | 3.5492043 | 4.3416257 | 65-69 |
| TTTTT | 18119865 | 3.1749835 | 4.242848 | 95-97 |
| CCCCC | 2074405 | 3.036577 | 12.608274 | 1 |
| GCAGG | 1976725 | 1.9669305 | 11.859514 | 4 |
| GGGGA | 1899915 | 1.9172294 | 5.8499823 | 15-19 |
| GGCCG | 1239100 | 1.8918583 | 9.288178 | 25-29 |
| CGGCC | 1237960 | 1.8637674 | 9.103401 | 25-29 |
| TGCAG | 2874805 | 1.8449011 | 8.169329 | 3 |
| CTGCA | 2872515 | 1.8177322 | 9.230598 | 2 |
| GGCCC | 1197850 | 1.8033812 | 34.447353 | 9 |
| CCCCA | 1854780 | 1.7694719 | 10.886289 | 1 |
| GCGGC | 1156995 | 1.7665005 | 8.738608 | 20-24 |
| CTGGG | 1717235 | 1.7148697 | 5.164394 | 1 |
| GCCGC | 1131500 | 1.7034903 | 8.869117 | 25-29 |
| CGGGG | 1053450 | 1.631148 | 8.304113 | 10-14 |
| TGGCC | 1645845 | 1.6206646 | 23.55618 | 8 |
| CCTCC | 1673385 | 1.6021595 | 5.4868855 | 1 |
| GCCCA | 1621055 | 1.5683615 | 5.1635866 | 1 |
| CGCGG | 1019780 | 1.5570005 | 8.386023 | 20-24 |
| CCTGG | 1580950 | 1.5567626 | 5.04937 | 1 |
| CAGGA | 2420445 | 1.5477517 | 7.44542 | 5 |
| TCTGC | 2411150 | 1.5312654 | 7.2380004 | 1 |
| CCCAG | 1577830 | 1.5265416 | 6.18334 | 1 |
| CTGGA | 2369995 | 1.5209402 | 18.118704 | 2 |
| GCCCC | 980320 | 1.4553108 | 8.310058 | 1 |
| CCGGG | 939475 | 1.4343908 | 7.6209726 | 10-14 |
| CCCGG | 946165 | 1.4244658 | 7.687745 | 10-14 |
| CCCCT | 1483780 | 1.4206249 | 8.321901 | 1 |
| CCCTG | 1447980 | 1.4059491 | 5.9500694 | 1 |
| ACTGG | 2190345 | 1.40565 | 16.909056 | 1 |
| CTCCA | 2251205 | 1.4047059 | 5.1463513 | 1 |
| ATGCA | 3328755 | 1.3728085 | 12.25127 | 6 |
| CTCCC | 1411690 | 1.3516033 | 7.008748 | 1 |
| ATGGC | 2105010 | 1.3508867 | 15.449282 | 7 |
| TGCAT | 3237420 | 1.3399411 | 11.609896 | 7 |
| GCCCG | 852230 | 1.283045 | 7.5077252 | 10-14 |
| CTCCT | 2045990 | 1.2812461 | 6.463061 | 1 |
| TCTGT | 2997975 | 1.2452984 | 5.1911607 | 9 |
| TCCTT | 2921100 | 1.1964501 | 6.1175714 | 3 |
| GCATC | 1841765 | 1.1654719 | 16.51944 | 8 |
| CCCCG | 784795 | 1.165049 | 7.1871552 | 1 |
| CATCT | 2849655 | 1.1630055 | 11.131097 | 9 |
| GATGC | 1800630 | 1.1555512 | 17.237217 | 5 |
| CCTAC | 1820780 | 1.1361295 | 6.230683 | 4 |
| GGATG | 1737570 | 1.1308478 | 17.781359 | 4 |
| GTTCC | 1750015 | 1.1113938 | 5.7071214 | 9 |
| GATCC | 1690835 | 1.069963 | 13.7700615 | 1 |
| TGGAT | 2329725 | 0.97788674 | 12.26007 | 3 |
| ATCCT | 2326565 | 0.9495214 | 8.483317 | 2 |
| TCGCG | 937640 | 0.9232947 | 5.670171 | 20-24 |
| TATGG | 2015135 | 0.8458397 | 9.684713 | 6 |
| CTACG | 1312405 | 0.830492 | 5.509676 | 5 |
| AGTAT | 2901945 | 0.7827739 | 6.4596767 | 4 |
| CGTTC | 1223375 | 0.776937 | 5.451186 | 8 |
| CCTTA | 1851760 | 0.75574315 | 5.071609 | 4 |
| GTATG | 1797520 | 0.7544971 | 9.852701 | 5 |
| TAGTA | 2378030 | 0.6414526 | 6.178138 | 3 |
| CTAGT | 1379905 | 0.5711312 | 9.524755 | 2 |
| ACTAG | 1339845 | 0.552564 | 9.382269 | 1 |