![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 3291_5903_10012_H94MGADXX_HK_CF70_GCCAAT_R1.fastq |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 11952919 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 38 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
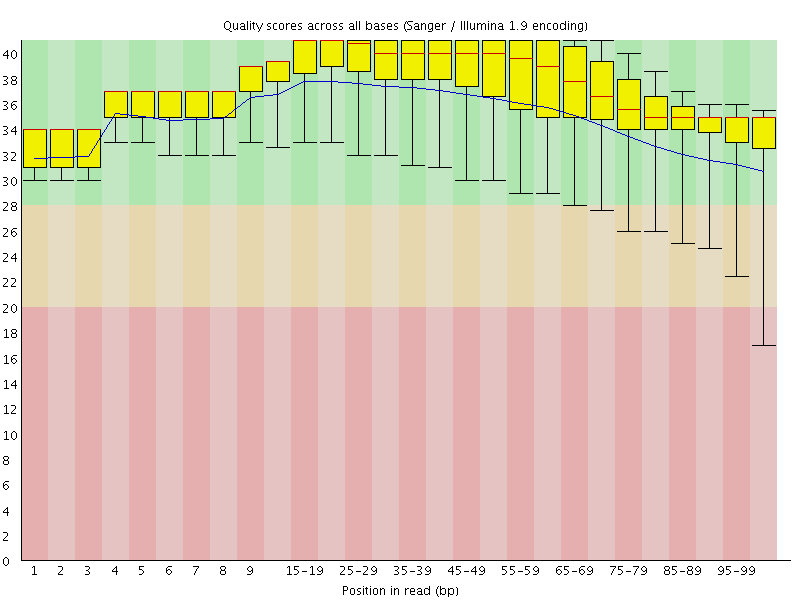
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
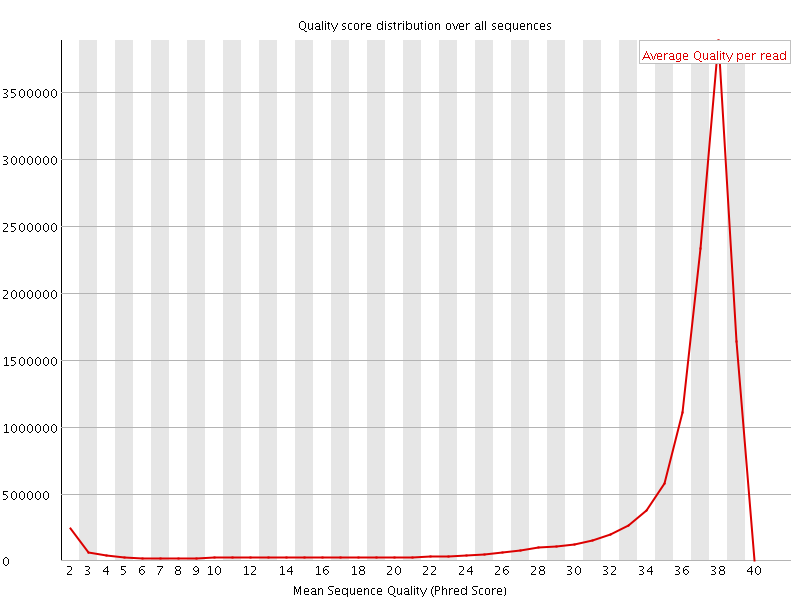
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
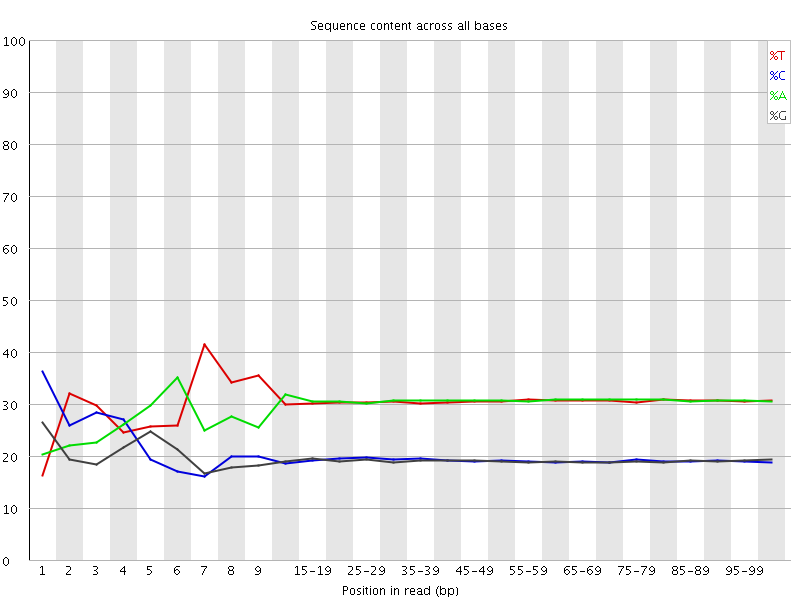
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
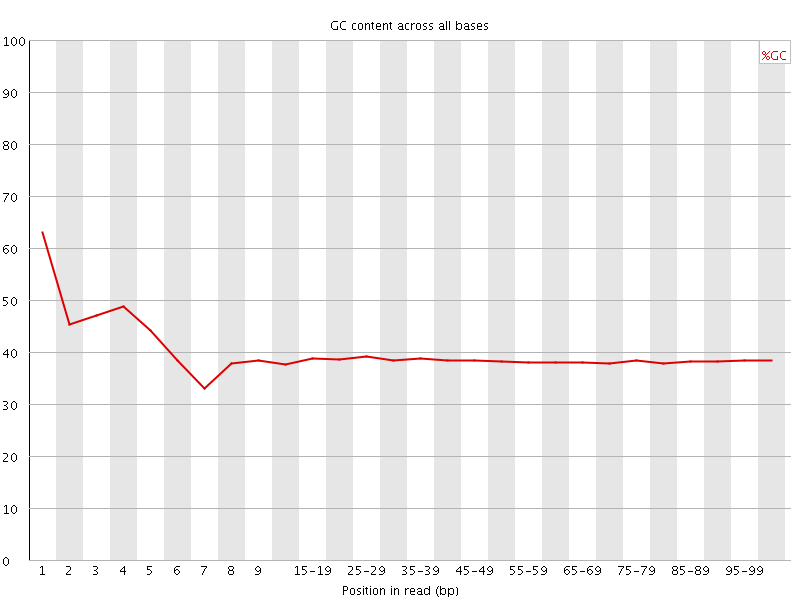
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
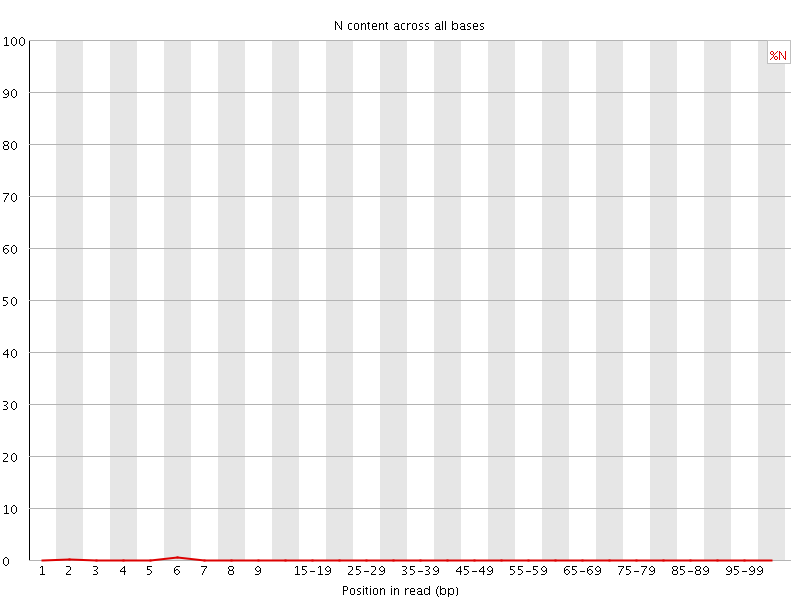
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
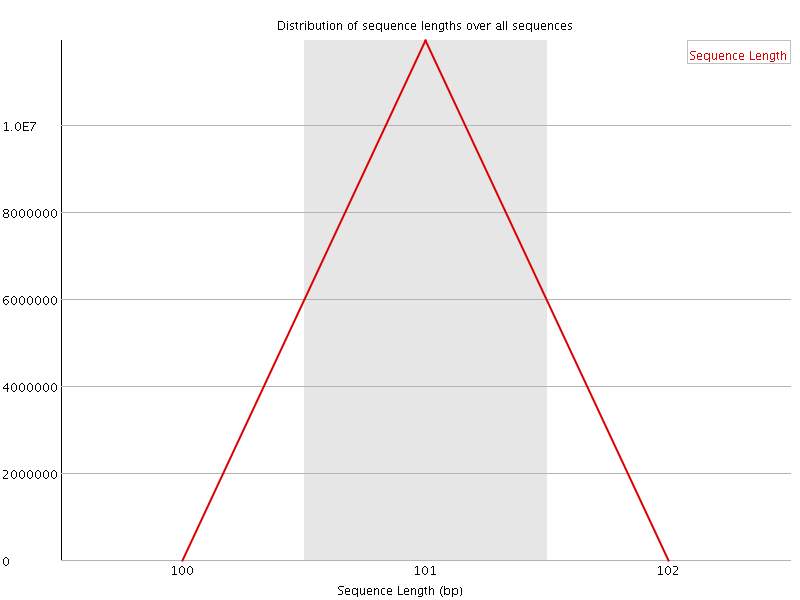
![[FAIL]](Icons/error.png) Sequence Duplication Levels
Sequence Duplication Levels
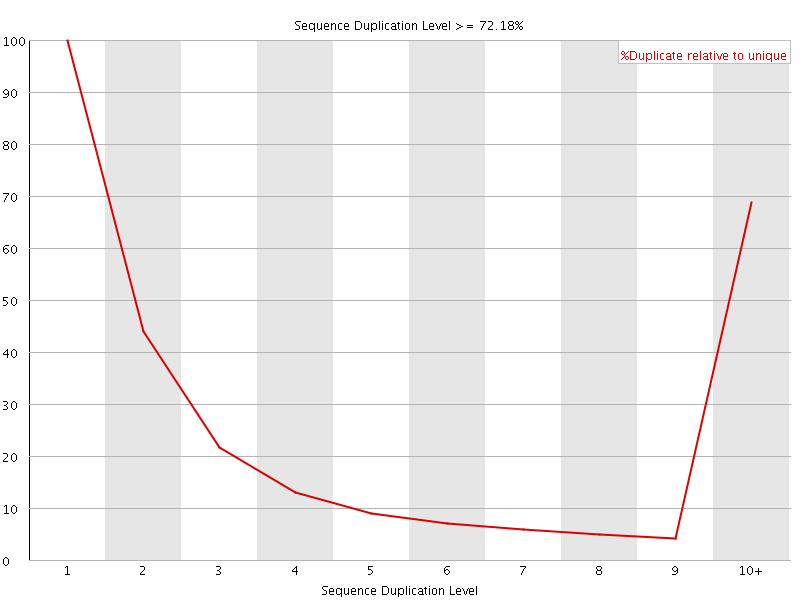
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| ACTAGTATGGCCCGGGGGATCCTACGTTCCAAATGCAGCGAGCTCGTATA | 32672 | 0.273339089807268 | No Hit |
| ACTGGATGCATCTGCAGGATATCGCGGCCGCTCGACGTAGAACTCAATCT | 28987 | 0.24250980032576144 | No Hit |
| GATCCTTATCTGTCAAAACCGCTAATGTCCGTTCTAAGACCGTCTGGAGA | 17944 | 0.1501223257682914 | No Hit |
| CTCCTGTTCACTTTTCTTCCCCAATTCGAGGCTCTACACGCAAATGGCTG | 17602 | 0.14726109998737547 | No Hit |
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGCCAATATCTCGTATGC | 17453 | 0.1460145425565086 | TruSeq Adapter, Index 6 (100% over 50bp) |
| TCTGCAGGATATCGCGGCCGCCATCTGCCCTACGTTTGAGGGTTATAAGC | 15369 | 0.12857947083888044 | No Hit |
| GATCCTACGTTCCAAATGCAGCGAGCTCGTATAACCCTTTAAGAGTTGCT | 15221 | 0.1273412795652677 | No Hit |
| ACTAGTATGGCCCGGGGGATCCTTATCTGTCAAAACCGCTAATGTCCGTT | 14291 | 0.11956075331891734 | No Hit |
| ACTGGATGCATCTGCAGGATATCGCGGCCGCCATCTGCCCTACGTTTGAG | 13584 | 0.11364588014024023 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
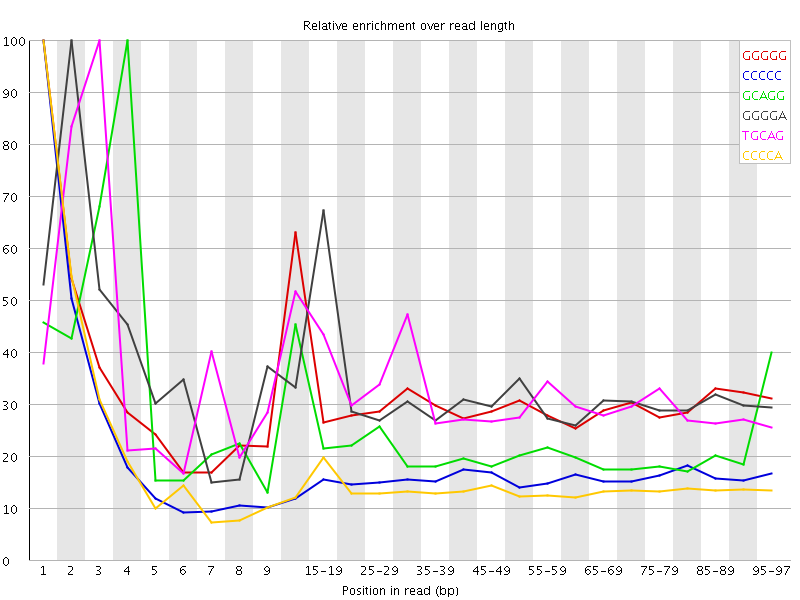
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GGGGG | 916520 | 2.9506269 | 9.314832 | 1 |
| CCCCC | 765985 | 2.2488577 | 13.391532 | 1 |
| GCAGG | 854390 | 1.710347 | 7.316632 | 4 |
| GGGGA | 813415 | 1.6586181 | 5.0348263 | 2 |
| TGCAG | 1303995 | 1.6493447 | 5.0010357 | 3 |
| CCCCA | 868020 | 1.6441424 | 10.977383 | 1 |
| GGCCC | 535845 | 1.6322744 | 17.113039 | 9 |
| CTGCA | 1301375 | 1.6159647 | 6.201997 | 2 |
| CCTGG | 776380 | 1.5220956 | 5.538975 | 1 |
| CCCAA | 1241155 | 1.516714 | 5.3738413 | 2 |
| GCCCA | 779910 | 1.504736 | 5.9257545 | 1 |
| TGGCC | 758965 | 1.4879533 | 12.107091 | 8 |
| CCTCC | 764200 | 1.4439838 | 6.1461368 | 1 |
| CCCAG | 736010 | 1.4200367 | 6.4595976 | 1 |
| CCTGC | 730945 | 1.4068446 | 5.0606174 | 1 |
| CCCCT | 743980 | 1.4057775 | 9.595471 | 1 |
| GCCCC | 469865 | 1.4051443 | 7.7568707 | 1 |
| CCCTG | 727835 | 1.4008589 | 6.781808 | 1 |
| CCAGT | 1125080 | 1.3970528 | 5.6718807 | 3 |
| CTGGA | 1093995 | 1.3837285 | 8.521821 | 2 |
| ACTGG | 1093620 | 1.3832542 | 7.230093 | 1 |
| ATGGC | 1081045 | 1.3673489 | 7.980058 | 7 |
| CTCCA | 1117060 | 1.3617575 | 5.670142 | 1 |
| CGGGG | 425910 | 1.346121 | 5.242442 | 1 |
| ATGCA | 1650125 | 1.3219476 | 6.1385746 | 6 |
| TGCAT | 1628500 | 1.3014598 | 5.2666054 | 7 |
| CCCAC | 674330 | 1.2772684 | 5.2998705 | 1 |
| CTCCC | 670895 | 1.2676806 | 8.243854 | 1 |
| CTCCT | 1011510 | 1.2300962 | 8.351829 | 1 |
| TCCTG | 983815 | 1.2186767 | 6.3403435 | 2 |
| CCTGT | 948980 | 1.1755258 | 6.837055 | 3 |
| CCCGG | 380705 | 1.1596917 | 5.7414746 | 1 |
| GATGC | 872050 | 1.1030037 | 7.5914335 | 5 |
| CCCCG | 368310 | 1.1014413 | 8.03172 | 1 |
| GCATC | 873055 | 1.0841043 | 6.841208 | 8 |
| GGATG | 835935 | 1.0769963 | 7.931936 | 4 |
| CCTCG | 494435 | 0.9516355 | 5.4939566 | 1 |
| CTCGG | 484800 | 0.950452 | 5.747759 | 1 |
| TGGAT | 1147760 | 0.93432987 | 5.727342 | 3 |
| GATCC | 732655 | 0.90976447 | 5.4761405 | 1 |
| CCCGC | 301325 | 0.9011209 | 5.355935 | 1 |