![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 3291_5903_10012_H94MGADXX_HK_CF70_GCCAAT_R2.fastq |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 11952919 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 38 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
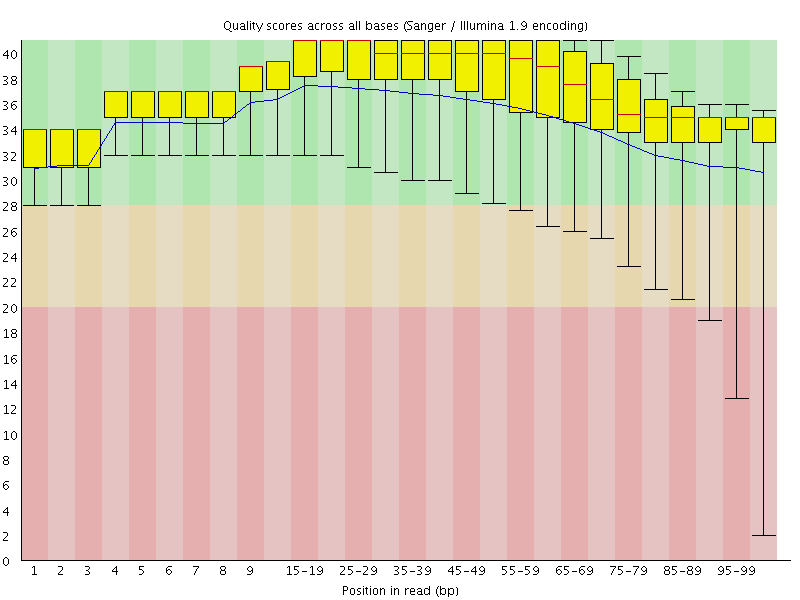
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
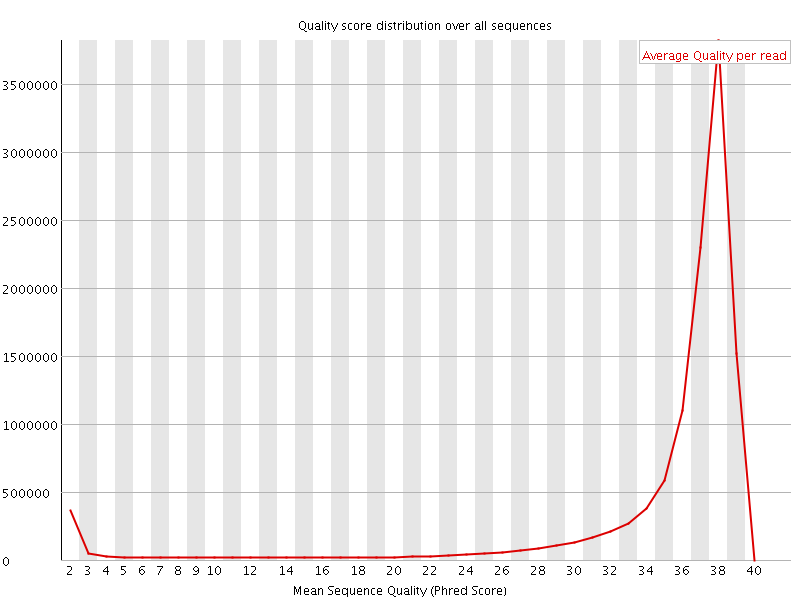
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
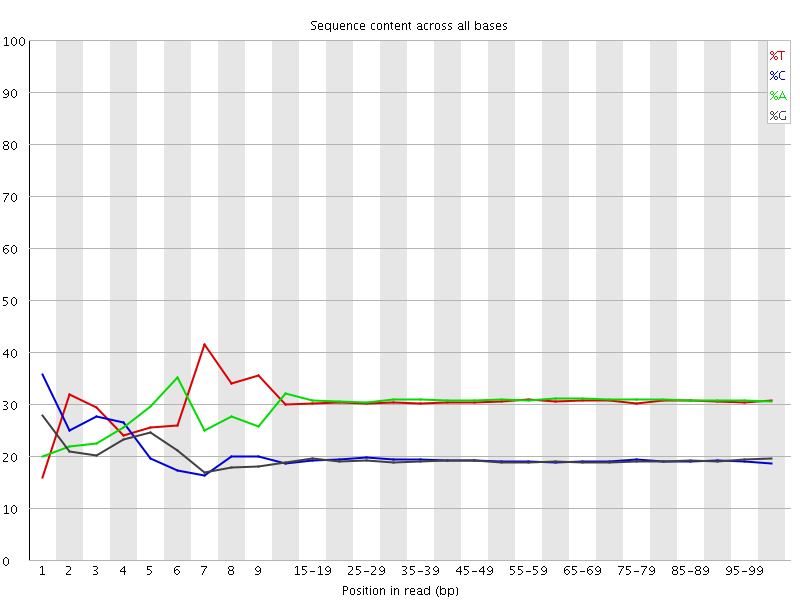
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
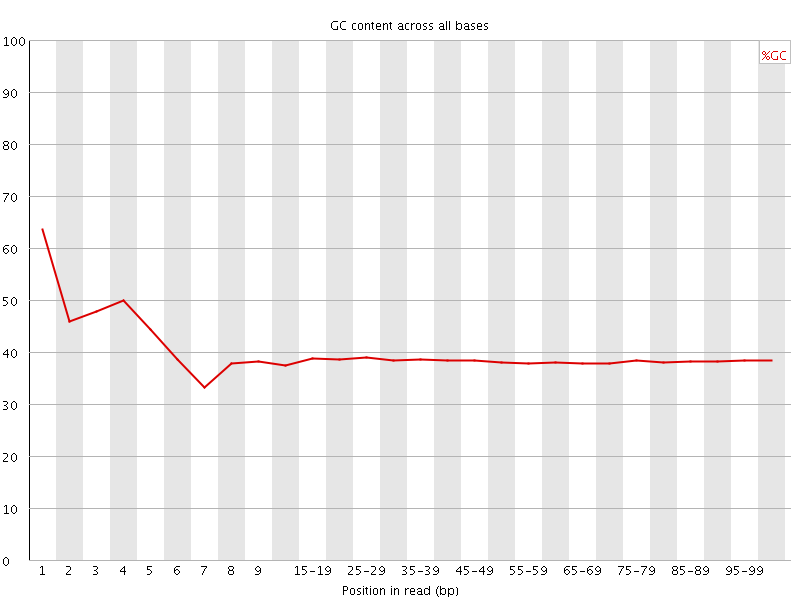
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
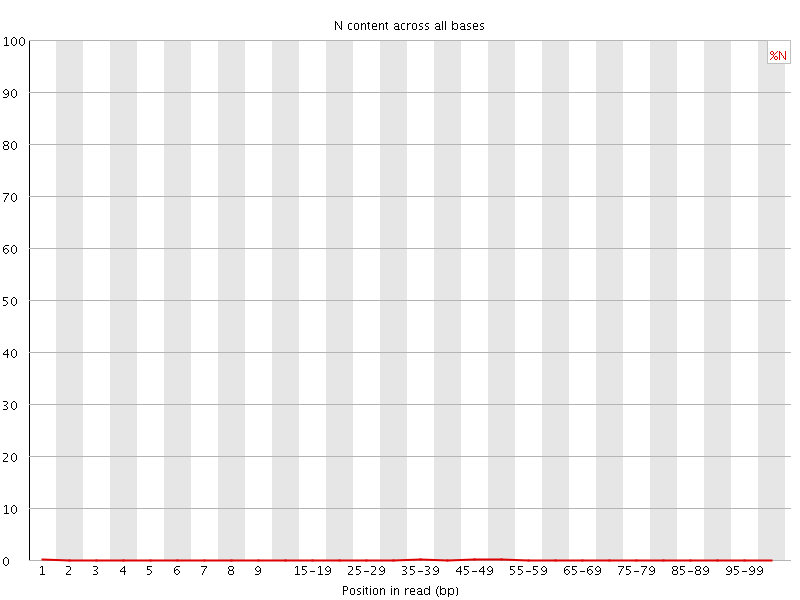
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
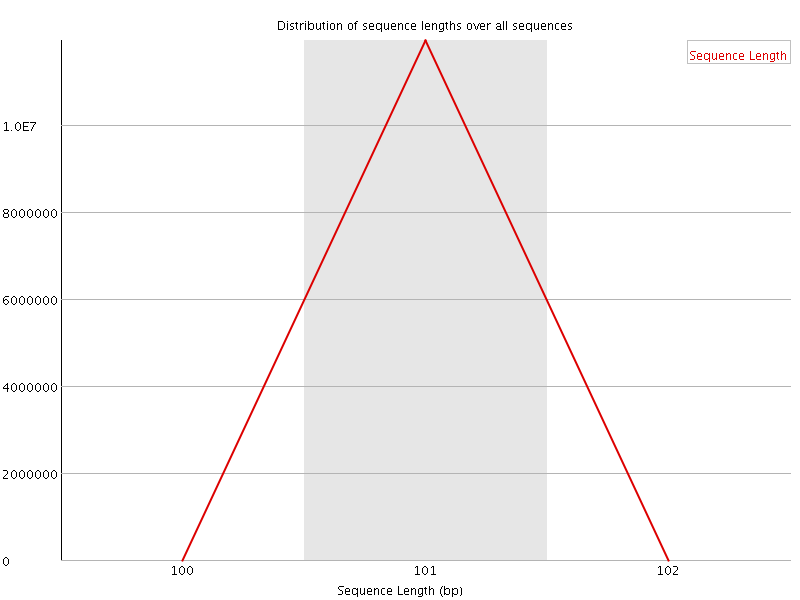
![[FAIL]](Icons/error.png) Sequence Duplication Levels
Sequence Duplication Levels
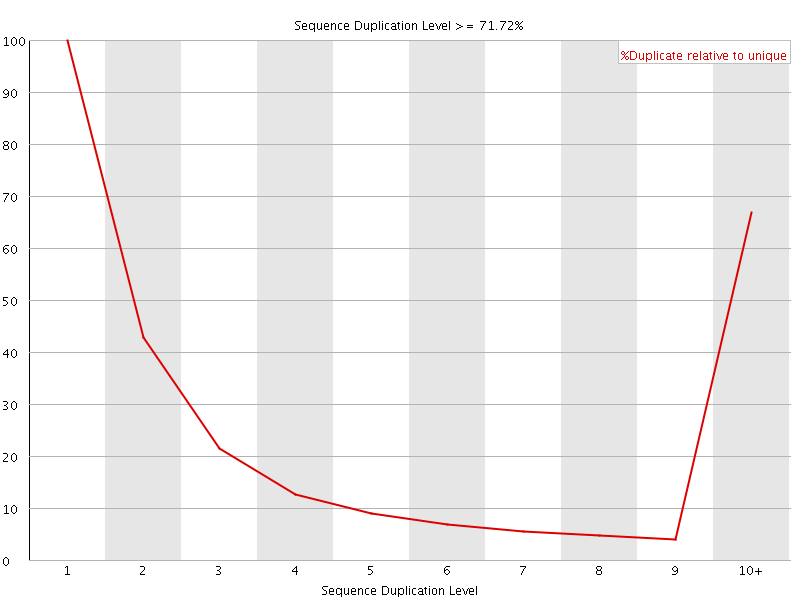
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| ACTGGATGCATCTGCAGGATATCGCGGCCGCTCGACGTAGAACTCAATCT | 32602 | 0.2727534587994782 | No Hit |
| ACTAGTATGGCCCGGGGGATCCTACGTTCCAAATGCAGCGAGCTCGTATA | 28397 | 0.23757376754581874 | No Hit |
| GATCCTTATCTGTCAAAACCGCTAATGTCCGTTCTAAGACCGTCTGGAGA | 22851 | 0.19117505941435728 | No Hit |
| CTCCTGTTCACTTTTCTTCCCCAATTCGAGGCTCTACACGCAAATGGCTG | 18355 | 0.153560816399743 | No Hit |
| GATCCTACGTTCCAAATGCAGCGAGCTCGTATAACCCTTTAAGAGTTGCT | 15827 | 0.13241117086127666 | No Hit |
| ACTGGATGCATCTGCAGGATATCGCGGCCGCCATCTGCCCTACGTTTGAG | 14383 | 0.12033043978629822 | No Hit |
| ACTAGTATGGCCCGGGGGATCCTTATCTGTCAAAACCGCTAATGTCCGTT | 13149 | 0.11000660173468924 | No Hit |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
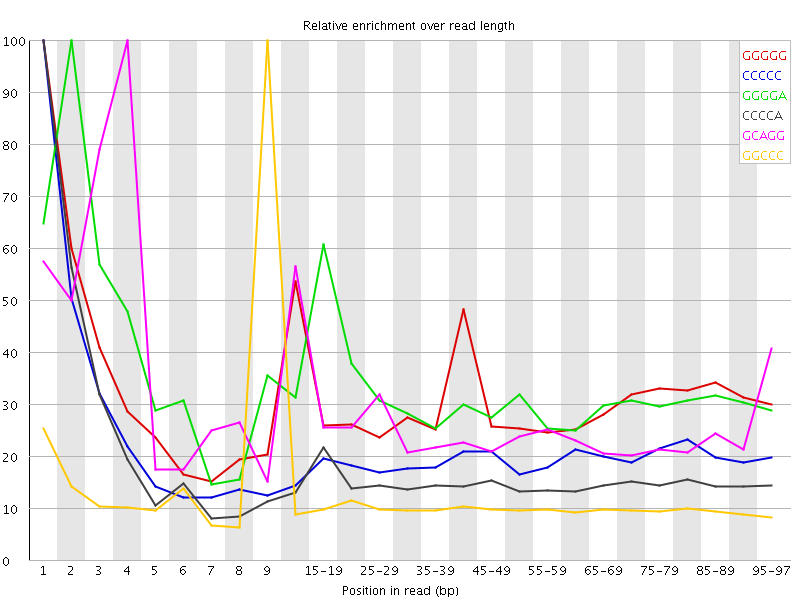
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GGGGG | 1002190 | 3.1787612 | 10.152218 | 1 |
| CCCCC | 925895 | 2.7677915 | 13.668476 | 1 |
| GGGGA | 858020 | 1.7247527 | 5.2676635 | 2 |
| CCCCA | 890850 | 1.7078315 | 10.626501 | 1 |
| GCAGG | 851495 | 1.69147 | 6.1855135 | 4 |
| GGCCC | 550050 | 1.6837139 | 15.469249 | 9 |
| CTGCA | 1291395 | 1.6114213 | 5.4970655 | 2 |
| GGGCC | 504400 | 1.5623864 | 5.121137 | 1 |
| CCCAA | 1250485 | 1.5374031 | 5.1402106 | 2 |
| CTGGG | 771120 | 1.5363765 | 5.339148 | 1 |
| CCTGG | 779105 | 1.5339968 | 5.61754 | 1 |
| GCCCA | 779785 | 1.512734 | 5.8616977 | 1 |
| TGGCC | 759480 | 1.4953567 | 11.003849 | 8 |
| GCCCC | 490940 | 1.4850714 | 8.097348 | 1 |
| CCCCT | 764285 | 1.4695666 | 9.25777 | 1 |
| CCTCC | 757890 | 1.4572703 | 5.776303 | 1 |
| CGGGG | 455025 | 1.4262508 | 6.0725265 | 1 |
| CCCAG | 734285 | 1.4244668 | 6.2036214 | 1 |
| CCCTG | 724890 | 1.4104357 | 6.0898614 | 1 |
| CCTGC | 720010 | 1.4009407 | 5.107321 | 1 |
| CTGGA | 1107695 | 1.3986771 | 9.063715 | 2 |
| ACTGG | 1089910 | 1.3762201 | 7.7998915 | 1 |
| CCAGT | 1092560 | 1.3633121 | 5.66583 | 3 |
| ATGGC | 1069380 | 1.3502971 | 7.1796665 | 7 |
| CTCCA | 1071175 | 1.3208797 | 5.486859 | 1 |
| ATGCA | 1637680 | 1.3105328 | 6.427608 | 6 |
| CTCCC | 677575 | 1.3028408 | 8.354027 | 1 |
| TGCAT | 1622540 | 1.3022902 | 5.638433 | 7 |
| CCCAC | 676985 | 1.297835 | 5.0395827 | 1 |
| CTCCT | 988245 | 1.2222525 | 8.121521 | 1 |
| TCCTG | 975245 | 1.2205547 | 6.1444044 | 2 |
| CCCGG | 389545 | 1.1924047 | 6.0358024 | 1 |
| CCTGT | 942675 | 1.1797922 | 6.816849 | 3 |
| CCCCG | 383675 | 1.1605995 | 7.683153 | 1 |
| GCATC | 878845 | 1.0966356 | 7.452653 | 8 |
| GATGC | 867155 | 1.0949492 | 8.098734 | 5 |
| CATCT | 1368830 | 1.0857121 | 5.2243366 | 9 |
| GGATG | 842320 | 1.0762709 | 8.503775 | 4 |
| CTCGG | 496080 | 0.97674274 | 5.7991824 | 1 |
| CCTCG | 488015 | 0.94954234 | 5.233917 | 1 |
| TGGAT | 1155245 | 0.9382826 | 6.12935 | 3 |
| CCCGC | 307025 | 0.9287367 | 5.3228345 | 1 |
| GATCC | 726270 | 0.90625024 | 6.515623 | 1 |