![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | BiGoRNA_GTGTCTAC_2.fastq |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 25542180 |
| Filtered Sequences | 0 |
| Sequence length | 50 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
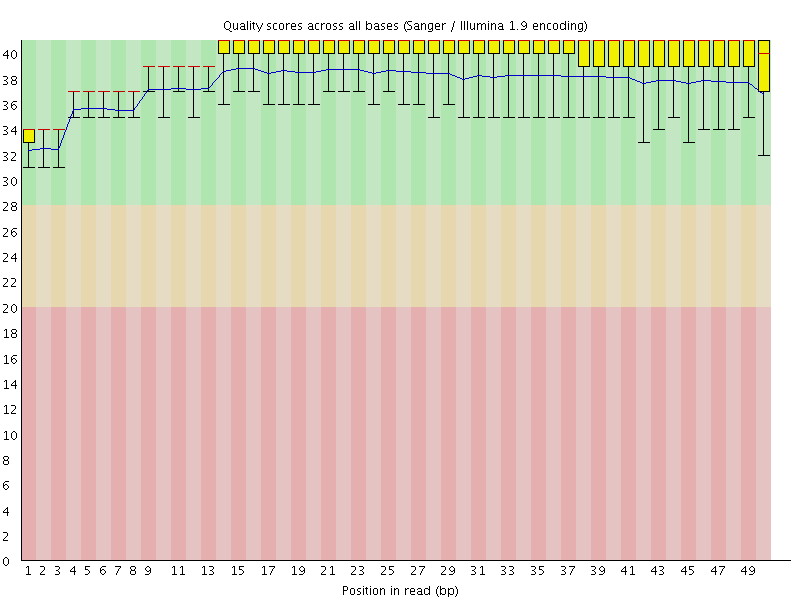
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
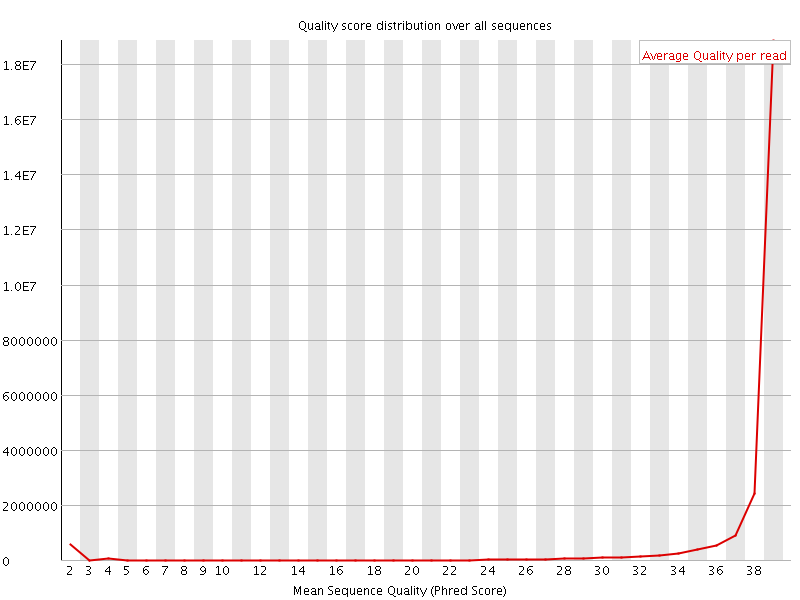
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
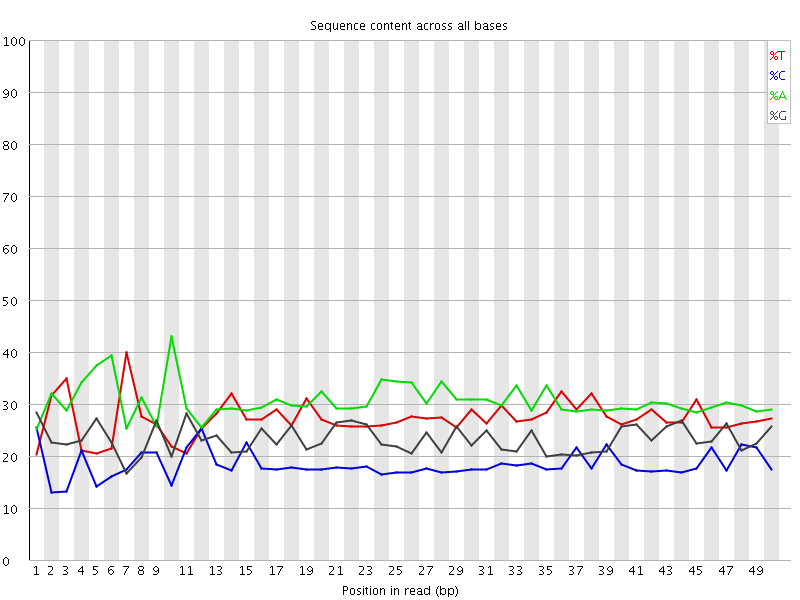
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
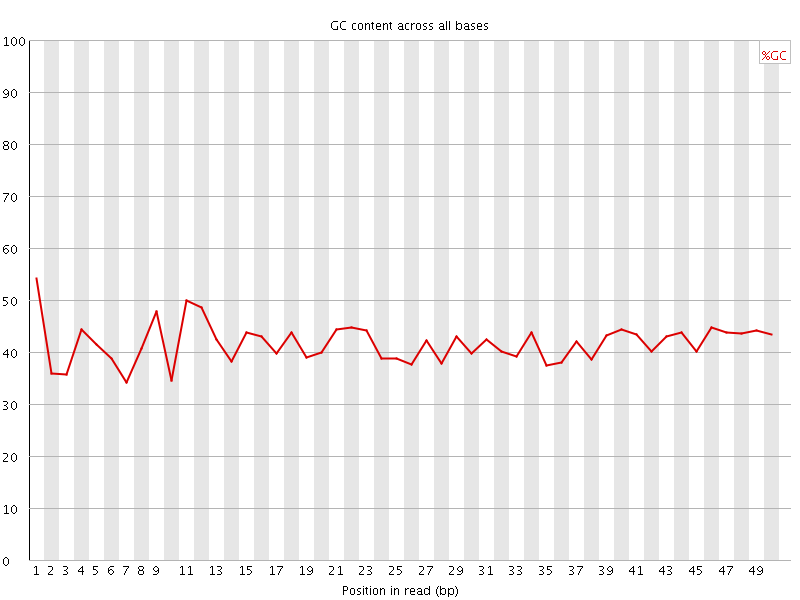
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
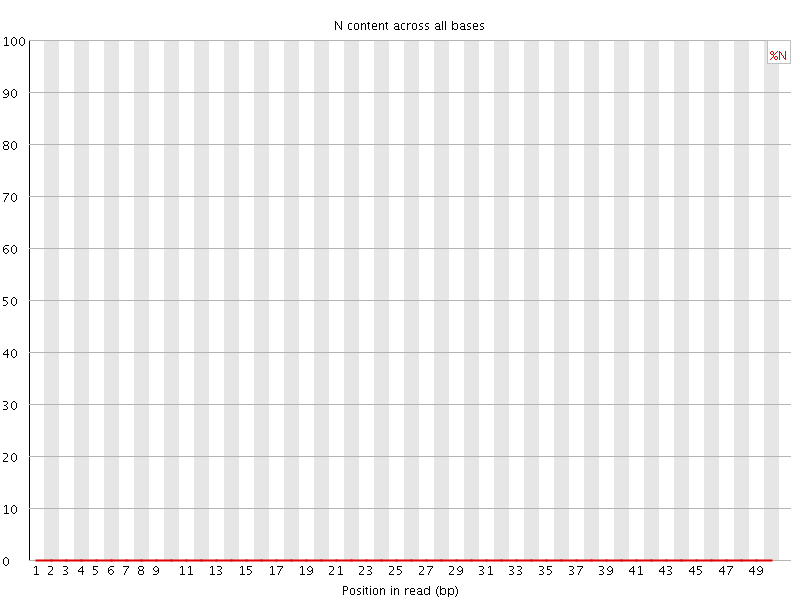
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
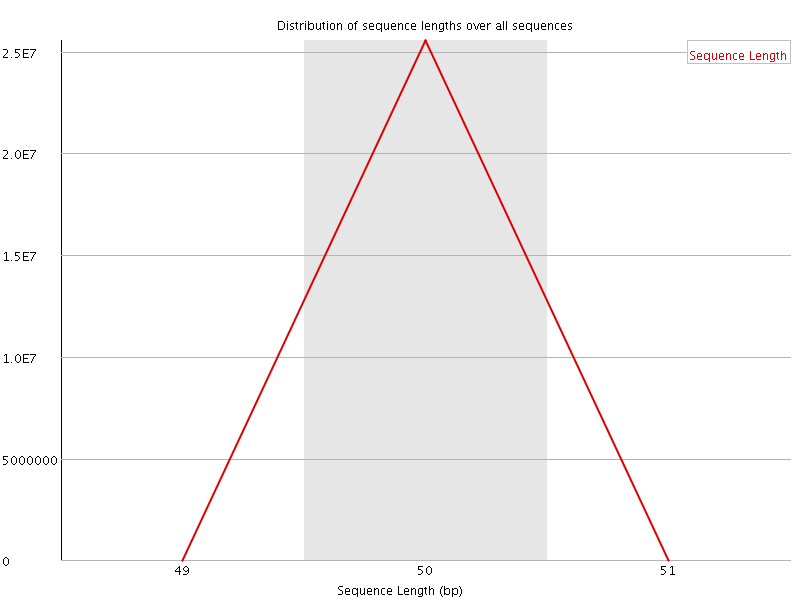
![[FAIL]](Icons/error.png) Sequence Duplication Levels
Sequence Duplication Levels
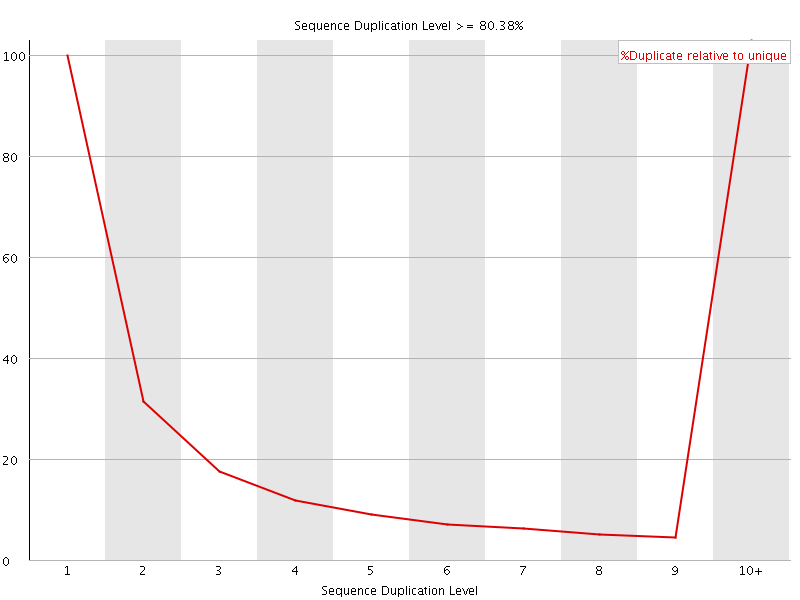
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCGTCGTGTAGGGAAAGAGTGTAGATCTCGGTGGTCGCCG | 564422 | 2.2097643975572954 | Illumina Single End PCR Primer 1 (100% over 50bp) |
| GGGGACTACTAGCAATAAATAGCAGTTTAAAAACTTTTTCTAAATGAAGT | 300638 | 1.1770256101867576 | No Hit |
| GGACTACTAGCAATAAATAGCAGTTTAAAAACTTTTTCTAAATGAAGTAT | 159005 | 0.6225192994489899 | No Hit |
| GGGACTACTAGCAATAAATAGCAGTTTAAAAACTTTTTCTAAATGAAGTA | 142359 | 0.5573486679680434 | No Hit |
| GATCGGAAGAGCGTCGTGTAGGGAAAGAGGGTAGATCTCGGTGGTCGCCG | 81870 | 0.32052863146372 | Illumina Single End PCR Primer 1 (98% over 50bp) |
| CTACTAGCAATAAATAGCAGTTTAAAAACTTTTTCTAAATGAAGTATAGG | 67912 | 0.26588176890147985 | No Hit |
| AATAAATAGCAGTTTAAAAACTTTTTCTAAATGAAGTATAGGAGCATTAG | 59011 | 0.23103352963607646 | No Hit |
| CTAGCAATAAATAGCAGTTTAAAAACTTTTTCTAAATGAAGTATAGGAGC | 56836 | 0.22251820322306085 | No Hit |
| TAGCAATAAATAGCAGTTTAAAAACTTTTTCTAAATGAAGTATAGGAGCA | 49995 | 0.19573505472124933 | No Hit |
| GACTACTAGCAATAAATAGCAGTTTAAAAACTTTTTCTAAATGAAGTATA | 46077 | 0.1803957219000101 | No Hit |
| GCAAAATTAAACCTCGCCTGTTTAACAAAAACATCACTAGAAGATAAAGA | 39880 | 0.15613389303497197 | No Hit |
| AAAGCATTGTTTTTAAGCGTTAAAAAAAAGTACCTTTAGTATAATGGCCT | 39797 | 0.15580894034886608 | No Hit |
| ACTAGCAATAAATAGCAGTTTAAAAACTTTTTCTAAATGAAGTATAGGAG | 39572 | 0.15492804451303688 | No Hit |
| TTTAAAAACTTTTTCTAAATGAAGTATAGGAGCATTAGAAACATTATTTT | 38759 | 0.15174507422624067 | No Hit |
| CTTTAATGTGGGGCGTAGTGTGTTAAAGATACTCACTAGACCAGGATGTT | 37058 | 0.1450855017073719 | No Hit |
| CTTGAATAAACTTTGCTAAGGATGTGATATGCAATTTCGAGTTTTAAGAT | 36215 | 0.14178507864246515 | No Hit |
| GCAATAAATAGCAGTTTAAAAACTTTTTCTAAATGAAGTATAGGAGCATT | 34945 | 0.13681291103578472 | No Hit |
| TTTCAAGAGATTTTGCTGTATTAGCACCTCCCGAAATAAAAAGATTTTTA | 34384 | 0.13461654408511725 | No Hit |
| ACTACTAGCAATAAATAGCAGTTTAAAAACTTTTTCTAAATGAAGTATAG | 32028 | 0.12539258591083455 | No Hit |
| TTATAAGTGAAAAATTCGGCTAAAGGTTGGATTTAAAATGCTTTCTAAGT | 31068 | 0.12163409701129661 | No Hit |
| GATCGGAAGAGCGTCGTGTAGGGAAAGAGTGTAGATCTCGGGGGTCGCCG | 30815 | 0.1206435785825642 | Illumina Single End PCR Primer 1 (98% over 50bp) |
| CGAAAAATGCTTTTAAGTGCAGCTTTAATGTGGGGCGTAGTGTGTTAAAG | 29802 | 0.11667758977503095 | No Hit |
| TAGAAACATTATTTTAGATTTTGTCTGGTAAGATGAAAGCATTGTTTTTA | 29562 | 0.11573796755014647 | No Hit |
| GCAATTTCGAGTTTTAAGATAGCTGGTTGTTCGAAAAATGCTTTTAAGTG | 29346 | 0.11489230754775043 | No Hit |
| ATTGTTTTTAAGCGTTAAAAAAAAGTACCTTTAGTATAATGGCCTTTCAA | 28961 | 0.11338499689533157 | No Hit |
| AAAAAATATTTAAAGGAACTCGGCAAAATTAAACCTCGCCTGTTTAACAA | 27987 | 0.10957169669934202 | No Hit |
| GATCGGAAGAGCGTCGTGTAGGGAAAGAGGGTAGATCTCGGGGGTCGCCG | 27629 | 0.10817009354722266 | Illumina Single End PCR Primer 1 (97% over 41bp) |
| TGAAAGCATTGTTTTTAAGCGTTAAAAAAAAGTACCTTTAGTATAATGGC | 26780 | 0.10484617992669382 | No Hit |
| GTTAAAGATACTCACTAGACCAGGATGTTATAAGTGAAAAATTCGGCTAA | 26751 | 0.10473264224118693 | No Hit |
| AAAATATTTAAAGGAACTCGGCAAAATTAAACCTCGCCTGTTTAACAAAA | 26307 | 0.10299434112515063 | No Hit |
| CCGAAATAAAAAGATTTTTAGGAAAGCAGAATAATAGAGATGTTCGATGT | 26202 | 0.10258325640176368 | No Hit |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
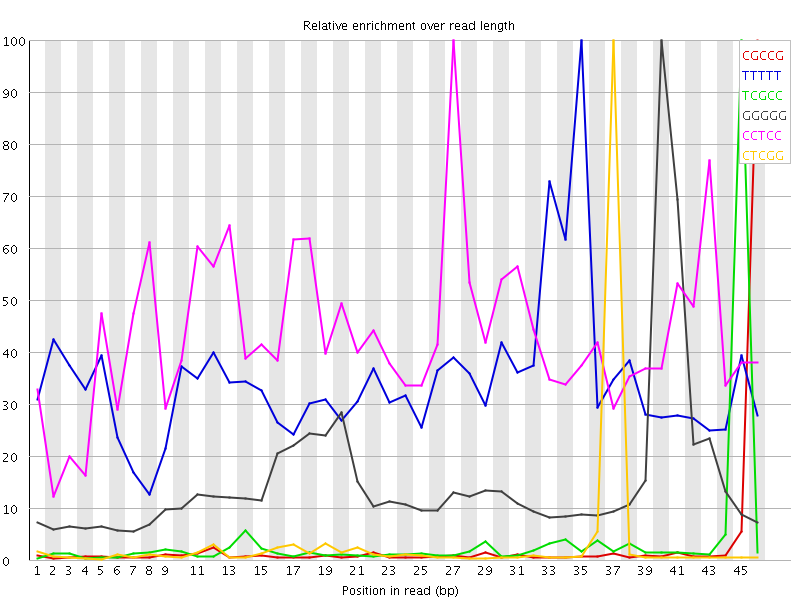
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CGCCG | 1595800 | 3.9286137 | 122.65764 | 46 |
| TTTTT | 7089195 | 3.9156537 | 11.306798 | 35 |
| TCGCC | 1644025 | 3.4607248 | 87.64958 | 45 |
| GGGGG | 2432045 | 2.9389355 | 19.073074 | 40 |
| CCTCC | 1037625 | 2.7689376 | 6.341659 | 27 |
| CTCGG | 1661985 | 2.7597616 | 82.206726 | 37 |
| CTTTT | 3221910 | 2.6383784 | 13.396796 | 34 |
| AAAAA | 8396910 | 2.6138613 | 6.6447377 | 29 |
| TTTCT | 3164660 | 2.5914974 | 13.575185 | 37 |
| AGCAG | 2852695 | 2.5401874 | 14.659685 | 21 |
| CTCCC | 942035 | 2.5138524 | 6.0770855 | 28 |
| AAGAG | 4675990 | 2.504141 | 28.768145 | 7 |
| TAGCA | 3109050 | 2.3672051 | 12.413114 | 10 |
| TTTTC | 2857585 | 2.3400378 | 13.268261 | 36 |
| GAAGA | 4351710 | 2.330479 | 29.117437 | 6 |
| CCACC | 975285 | 2.320574 | 5.123802 | 8 |
| TTTAA | 5256315 | 2.308187 | 7.465813 | 26 |
| ACTTT | 3033630 | 2.2150192 | 12.038235 | 33 |
| CGTCG | 1324920 | 2.2000582 | 75.9973 | 12 |
| ACCTC | 1361000 | 2.1842666 | 5.5729094 | 26 |
| TGAAG | 3594920 | 2.1591525 | 9.969974 | 45 |
| GAAAG | 3915020 | 2.0966175 | 25.743599 | 23 |
| ATCTC | 1933210 | 2.0927157 | 52.932713 | 35 |
| AAAGA | 5125015 | 2.0925183 | 19.53878 | 24 |
| GCGTC | 1255075 | 2.084079 | 75.51261 | 11 |
| GGTGG | 1990925 | 2.0571744 | 41.249897 | 40 |
| GGAAG | 2896150 | 2.034311 | 37.452785 | 5 |
| GTCGC | 1224285 | 2.0329516 | 68.81541 | 44 |
| TCTCG | 1394875 | 1.9805124 | 67.77008 | 36 |
| GCAGT | 1974850 | 1.9722112 | 15.407587 | 22 |
| AACTT | 3009605 | 1.9593657 | 10.800413 | 32 |
| CAGTT | 2290855 | 1.9562063 | 13.462169 | 23 |
| GGAAA | 3559920 | 1.9064502 | 26.071594 | 22 |
| TGGTC | 1690320 | 1.8932025 | 40.33111 | 42 |
| GGGGC | 1227175 | 1.8799188 | 13.732955 | 42 |
| AGAGC | 2069770 | 1.8430305 | 46.57932 | 8 |
| AGCGT | 1831665 | 1.8292174 | 45.954197 | 10 |
| TTCTA | 2500060 | 1.8254303 | 11.477643 | 38 |
| GAGGG | 1980395 | 1.8245645 | 17.07997 | 27 |
| CTAGC | 1415490 | 1.7920092 | 19.044043 | 9 |
| ATGAA | 3907300 | 1.7892035 | 7.904406 | 44 |
| CCTCG | 838160 | 1.7643534 | 5.415825 | 12 |
| GGTCG | 1345535 | 1.7624849 | 54.555737 | 43 |
| CGGTG | 1345425 | 1.7623409 | 51.15019 | 39 |
| TTAAA | 4500935 | 1.7623148 | 7.1852317 | 27 |
| TCTAA | 2704275 | 1.7605842 | 10.353585 | 39 |
| AAATG | 3828615 | 1.7531729 | 7.6928296 | 42 |
| AAAAC | 3365890 | 1.7421604 | 8.644822 | 30 |
| GAGCG | 1485755 | 1.7352765 | 59.9152 | 9 |
| AGGGA | 2468050 | 1.7336054 | 32.866924 | 20 |
| AAACT | 2977535 | 1.7284387 | 9.686315 | 31 |
| CTAAA | 2955530 | 1.715665 | 9.399419 | 40 |
| GATCT | 1998535 | 1.7065886 | 41.743076 | 34 |
| GGGGA | 1841305 | 1.6964189 | 14.440153 | 1 |
| GTTTA | 2919520 | 1.6815592 | 9.254684 | 25 |
| GGGAA | 2385565 | 1.6756663 | 33.524494 | 21 |
| GGAGG | 1799060 | 1.6574981 | 5.934845 | 18 |
| GTGTA | 2446805 | 1.6481711 | 28.78809 | 16 |
| GAAGT | 2733445 | 1.6417401 | 9.552267 | 46 |
| CGCCC | 524955 | 1.6383137 | 7.499716 | 46 |
| TCGGA | 1632225 | 1.630044 | 52.26804 | 3 |
| GGGCG | 1056235 | 1.6180545 | 17.875359 | 43 |
| AGAGG | 2297790 | 1.6140115 | 13.216188 | 26 |
| CGTGT | 1437795 | 1.6103681 | 48.13593 | 15 |
| GCAAT | 2098605 | 1.5978608 | 12.0772505 | 12 |
| TAAAT | 4064925 | 1.5915977 | 6.9284697 | 16 |
| AGCAA | 2336770 | 1.5864094 | 10.942164 | 11 |
| AGATC | 2067240 | 1.5739799 | 37.18802 | 33 |
| ATCGG | 1568235 | 1.5661395 | 54.639297 | 2 |
| CGGAA | 1757790 | 1.5652274 | 47.14552 | 4 |
| AGTTT | 2714935 | 1.5637243 | 9.25395 | 24 |
| GTCGT | 1395710 | 1.5632316 | 47.178303 | 13 |
| TGTAG | 2309690 | 1.5558102 | 28.703114 | 17 |
| CGGGG | 1014890 | 1.5547178 | 20.732662 | 39 |
| TCGTG | 1367835 | 1.5320108 | 46.987694 | 14 |
| GATCG | 1526515 | 1.5244753 | 54.664032 | 1 |
| CGCCT | 722825 | 1.5215695 | 5.243436 | 15 |
| GTGGT | 1692955 | 1.4957514 | 32.242386 | 41 |
| CTACT | 1377390 | 1.491036 | 16.505934 | 6 |
| GCGCC | 593335 | 1.4606993 | 25.344564 | 45 |
| ATAGC | 1916640 | 1.459314 | 12.525346 | 19 |
| TAAAA | 4110120 | 1.4349157 | 6.0868187 | 28 |
| GGAGA | 2009600 | 1.4115814 | 6.582195 | 31 |
| ACTAC | 1428265 | 1.3785774 | 15.3758135 | 5 |
| CTTTA | 1874115 | 1.3683937 | 5.005464 | 1 |
| GGACT | 1336375 | 1.3345892 | 15.905746 | 3 |
| AGTGT | 1966535 | 1.3246605 | 22.682945 | 28 |
| TCGGT | 1180335 | 1.322006 | 43.39395 | 38 |
| AATGA | 2859070 | 1.3092054 | 7.4543843 | 43 |
| GGGAC | 1103505 | 1.2888306 | 18.007717 | 2 |
| ACTAG | 1684335 | 1.2824389 | 11.467359 | 8 |
| GTAGA | 2074205 | 1.2457925 | 27.638407 | 31 |
| GAGTG | 1529970 | 1.2052804 | 26.670607 | 27 |
| AGAGT | 1966125 | 1.1808785 | 20.544754 | 26 |
| AGGGG | 1280115 | 1.1793871 | 7.749598 | 20 |
| AAATA | 3351115 | 1.1699337 | 5.713176 | 17 |
| GGGAG | 1266385 | 1.1667376 | 6.868337 | 30 |
| GGCGC | 596180 | 1.157775 | 19.960775 | 44 |
| TAGGG | 1438620 | 1.1333165 | 37.813305 | 19 |
| GTAGG | 1434770 | 1.1302836 | 37.935387 | 18 |
| AATAA | 3167960 | 1.105991 | 5.697591 | 14 |
| CAATA | 1893025 | 1.0988882 | 9.1159115 | 13 |
| ATAAA | 3115240 | 1.0875854 | 5.654975 | 15 |
| AGGGT | 1342495 | 1.0575912 | 10.645992 | 28 |
| TAGAT | 2021655 | 1.0382427 | 23.14901 | 32 |
| AATAG | 2114720 | 0.9683579 | 7.218642 | 18 |
| TCGGG | 721330 | 0.94485325 | 16.93215 | 38 |
| GGCGG | 616670 | 0.9446815 | 9.5409975 | 13 |
| TACTA | 1393060 | 0.9069343 | 10.029182 | 7 |
| TGGGC | 672215 | 0.8805186 | 5.318015 | 42 |
| GGGTC | 671730 | 0.8798834 | 8.879125 | 42 |
| GCGGG | 572145 | 0.8764733 | 9.567563 | 14 |
| GACTA | 1150130 | 0.8756996 | 11.789883 | 4 |
| GGGGT | 791490 | 0.8178274 | 7.5689034 | 41 |
| CGGCG | 418895 | 0.8134895 | 11.774427 | 12 |
| GGTAG | 955850 | 0.7529999 | 10.012338 | 30 |
| GCGGC | 376600 | 0.73135304 | 11.80545 | 11 |
| GGGTA | 867895 | 0.6837107 | 10.067652 | 29 |
| GTCGG | 465070 | 0.6091843 | 6.1441436 | 13 |
| AGCGG | 482895 | 0.56399363 | 7.4702873 | 10 |
| CGGGT | 392430 | 0.51403487 | 8.640686 | 15 |