![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | fastq_quality_filter_out.fastq |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 47749573 |
| Filtered Sequences | 0 |
| Sequence length | 50 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
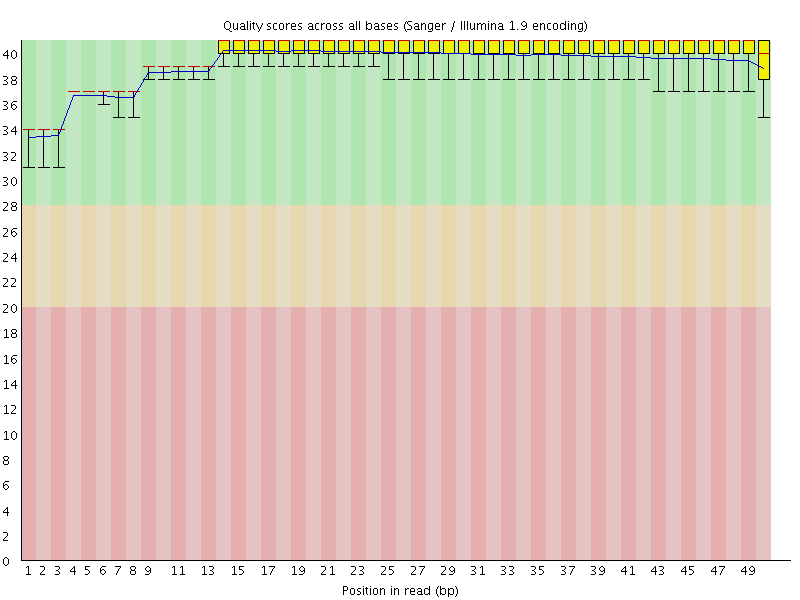
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
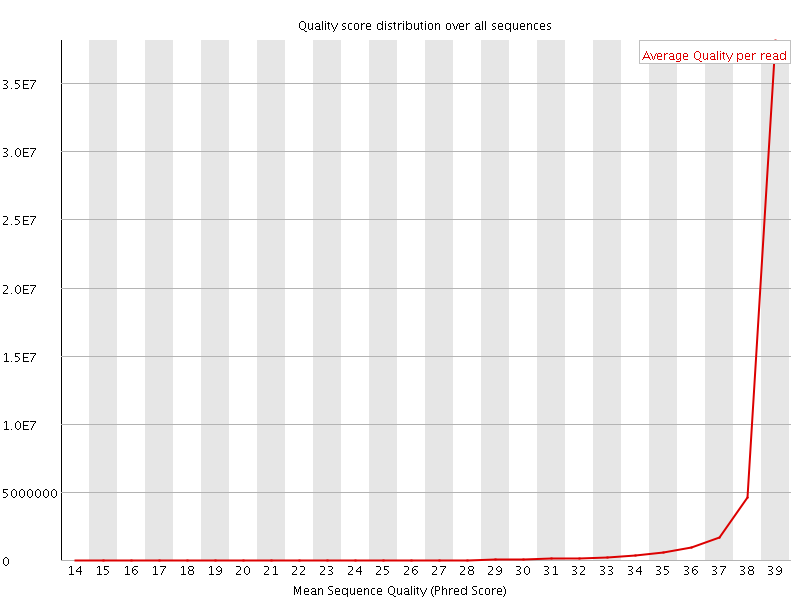
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
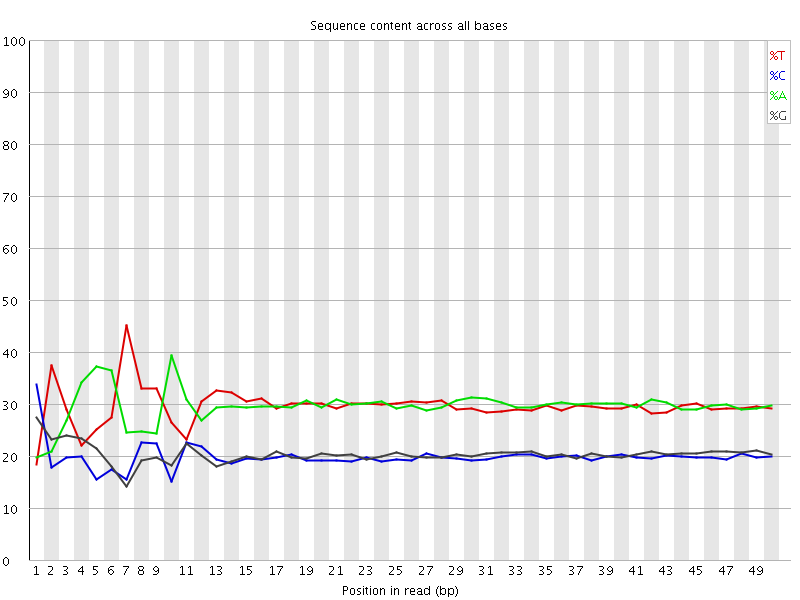
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
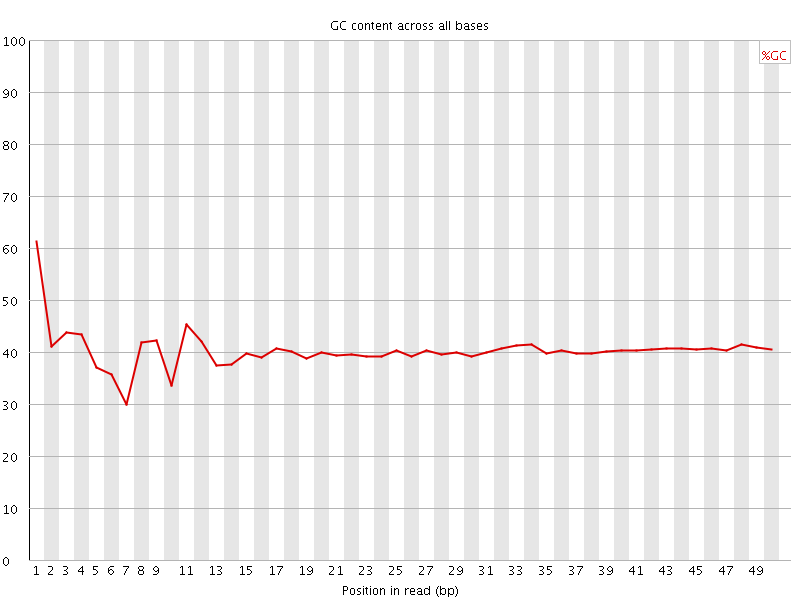
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
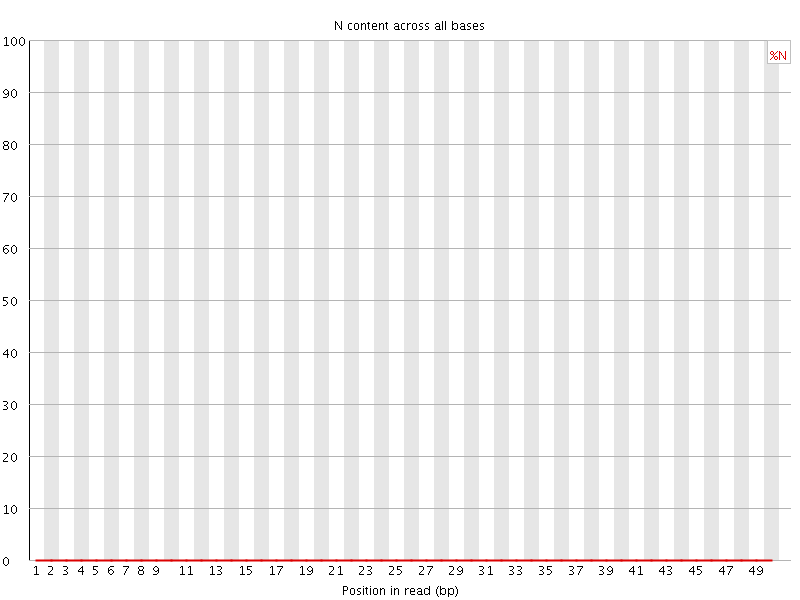
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
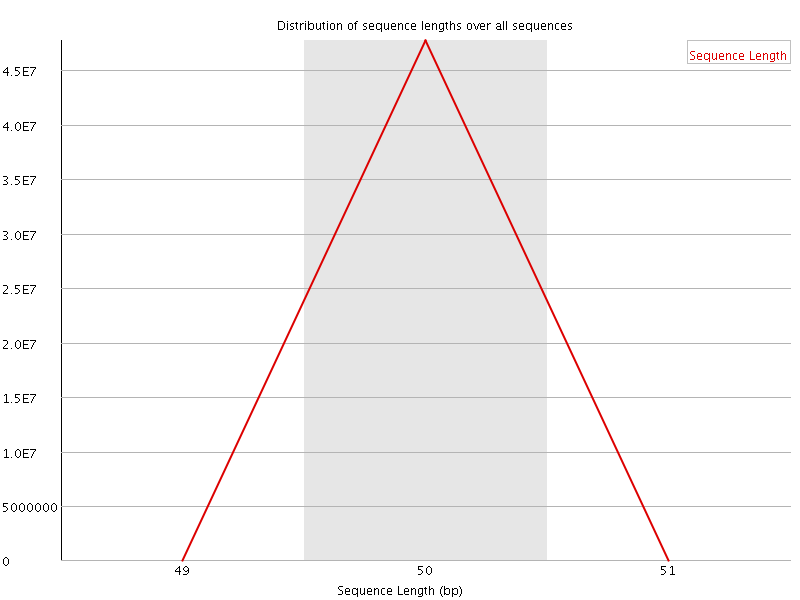
![[FAIL]](Icons/error.png) Sequence Duplication Levels
Sequence Duplication Levels
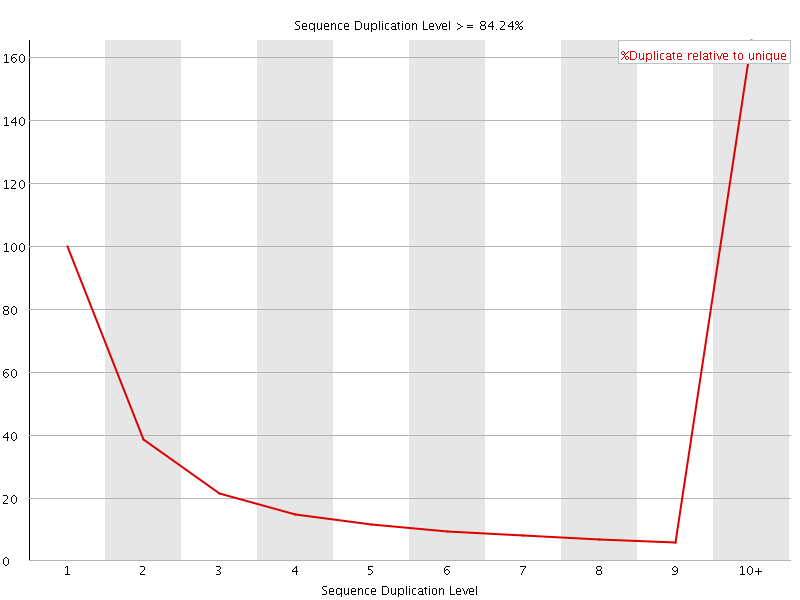
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GGGGACTACTAGCAATAAATAGCAGTTTAAAAACTTTTTCTAAATGAAGT | 300198 | 0.6286925330201383 | No Hit |
| GGACTACTAGCAATAAATAGCAGTTTAAAAACTTTTTCTAAATGAAGTAT | 158824 | 0.33261868121836397 | No Hit |
| ATAATAATCAATCAAAGAATATCTTAGACAGGTGAACGATTTGCTAAGCC | 151789 | 0.31788556517562994 | No Hit |
| GGGACTACTAGCAATAAATAGCAGTTTAAAAACTTTTTCTAAATGAAGTA | 142154 | 0.2977073742628023 | No Hit |
| GGCCATTATACTAAAGGTACTTTTTTTTAACGCTTAAAAACAATGCTTTC | 135326 | 0.2834077699501103 | No Hit |
| GGGAGGTGCTAATACAGCAAAATCTCTTGAAAGGCCATTATACTAAAGGT | 128931 | 0.2700149800292455 | No Hit |
| CAGCATAATAATCAATCAAAGAATATCTTAGACAGGTGAACGATTTGCTA | 125286 | 0.26238140391328735 | No Hit |
| CCCACATTAAAGCTGCACTTAAAAGCATTTTTCGAACAACCAGCTATCTT | 106223 | 0.2224585338176741 | No Hit |
| GTCCTTTCGTACATGCCGAGATAAGTTTATGTAAAGCAGATAGAAACCAA | 97903 | 0.2050342942333746 | No Hit |
| CCACATTAAAGCTGCACTTAAAAGCATTTTTCGAACAACCAGCTATCTTA | 90511 | 0.18955352752578541 | No Hit |
| ATTAAATTATCATTCTGCAATTTTTTTCTGGATTTACTCTTTCCAGTCTC | 88903 | 0.18618595814458905 | No Hit |
| CTAGTGAGTATCTTTAACACACTACGCCCCACATTAAAGCTGCACTTAAA | 82467 | 0.172707303581542 | No Hit |
| ATTTTTTAGAAATTATTCTTTTACTATTAGAAAGACAAGTTGTTAAATAC | 80417 | 0.1684140714724297 | No Hit |
| GAACGATTTGCTAAGCCTCATTTTTTAGAAATTATTCTTTTACTATTAGA | 77135 | 0.16154071157871924 | No Hit |
| CCACAATCAAAAGGCAAGGAATTTCGCTACCTTAGCACCCTCACGCTAGG | 68106 | 0.1426316419625365 | No Hit |
| CTACTAGCAATAAATAGCAGTTTAAAAACTTTTTCTAAATGAAGTATAGG | 67826 | 0.14204524928421872 | No Hit |
| AATAAATAGCAGTTTAAAAACTTTTTCTAAATGAAGTATAGGAGCATTAG | 58940 | 0.12343565878589113 | No Hit |
| GTCAAATCCGGAAAGAGTTAGTTATTCAGAAAGGTTAGACTTGCTTTCTA | 57917 | 0.12129323125046583 | No Hit |
| CTAGCAATAAATAGCAGTTTAAAAACTTTTTCTAAATGAAGTATAGGAGC | 56768 | 0.11888692700979756 | No Hit |
| CCAGCATAATAATCAATCAAAGAATATCTTAGACAGGTGAACGATTTGCT | 55798 | 0.11685549523133955 | No Hit |
| CTTAAAACTCGAAATTGCATATCACATCCTTAGCAAAGTTTATTCAAGAA | 55557 | 0.11635077867607319 | No Hit |
| CTCATTTTTTAGAAATTATTCTTTTACTATTAGAAAGACAAGTTGTTAAA | 55354 | 0.1159256439842928 | No Hit |
| CTTTCCTAAAAATCTTTTTATTTCGGGAGGTGCTAATACAGCAAAATCTC | 54185 | 0.1134774545523161 | No Hit |
| CCTGATTCAACATCGAGGTGCCAATCCCTTTAGCCAATACGAACTCTACT | 52180 | 0.10927846412364776 | No Hit |
| ATTATACTAAAGGTACTTTTTTTTAACGCTTAAAAACAATGCTTTCATCT | 51663 | 0.1081957319283253 | No Hit |
| GGAGGTGCTAATACAGCAAAATCTCTTGAAAGGCCATTATACTAAAGGTA | 51547 | 0.10795279781873651 | No Hit |
| CTGCTTTCCTAAAAATCTTTTTATTTCGGGAGGTGCTAATACAGCAAAAT | 50283 | 0.10530565372804486 | No Hit |
| TAGCAATAAATAGCAGTTTAAAAACTTTTTCTAAATGAAGTATAGGAGCA | 49916 | 0.10453706046753548 | No Hit |
| CTAAAAATCTTTTTATTTCGGGAGGTGCTAATACAGCAAAATCTCTTGAA | 48940 | 0.10249306313168498 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
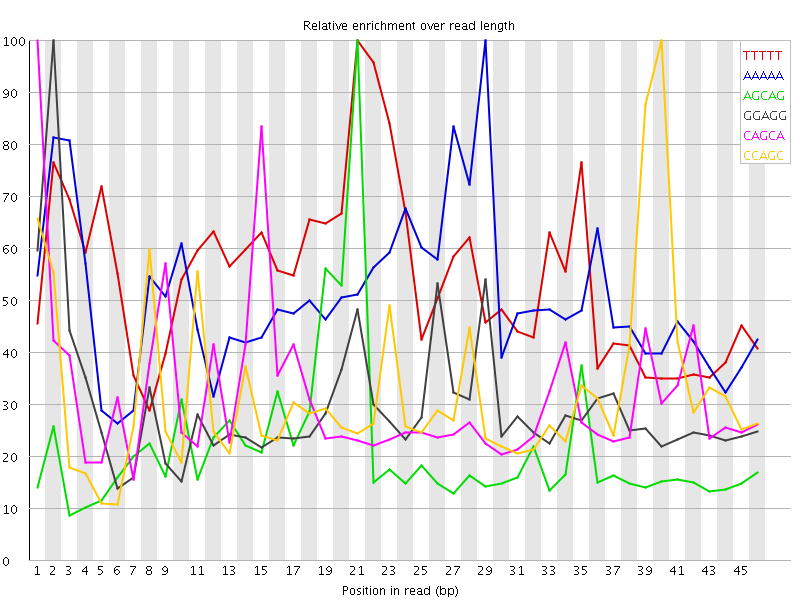
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 19774120 | 3.8650827 | 7.0923777 | 21 |
| AAAAA | 13824470 | 2.698305 | 5.3238044 | 29 |
| AGCAG | 4076185 | 2.4938018 | 11.579457 | 21 |
| GGAGG | 2874570 | 2.489291 | 8.300537 | 2 |
| CAGCA | 3800895 | 2.3843186 | 7.50487 | 1 |
| CCAGC | 2467170 | 2.3031006 | 7.056272 | 40 |
| GCTGC | 2381500 | 2.1687918 | 7.1271963 | 12 |
| CTTTT | 7346865 | 2.136364 | 5.521807 | 34 |
| TTTCT | 7344080 | 2.135554 | 5.8702555 | 37 |
| CTTTC | 4910330 | 2.1242003 | 5.8521504 | 4 |
| GAAAG | 4930385 | 2.0269988 | 5.186816 | 29 |
| TGAAG | 4792875 | 1.9710269 | 7.2995367 | 45 |
| TTTTC | 6767385 | 1.9678596 | 5.5676208 | 36 |
| GCCCC | 1411750 | 1.9611329 | 6.8153515 | 26 |
| AGGTG | 3258705 | 1.9449416 | 6.2915573 | 34 |
| CAGCT | 3019455 | 1.8946577 | 5.290381 | 40 |
| CCCCA | 1967695 | 1.8833973 | 5.1714644 | 27 |
| CTGCA | 2987685 | 1.8747227 | 5.6087537 | 13 |
| GGCCA | 2052985 | 1.8690859 | 9.323628 | 1 |
| AGCTG | 3016830 | 1.8462169 | 5.8432474 | 10 |
| AGCAA | 4259200 | 1.7954404 | 7.20144 | 11 |
| GGTGC | 2020945 | 1.7949462 | 7.270715 | 5 |
| GGGAG | 2067335 | 1.7902498 | 8.684742 | 1 |
| AAAGG | 4348945 | 1.7879547 | 5.0919733 | 30 |
| CTGGT | 2916725 | 1.7854643 | 5.760015 | 1 |
| CCCAC | 1860195 | 1.7805029 | 9.477313 | 1 |
| TAGCA | 4160910 | 1.7545072 | 6.9987626 | 10 |
| CCACA | 2724890 | 1.7526602 | 6.776251 | 1 |
| AAAAC | 5867375 | 1.7042047 | 6.0041885 | 30 |
| ACCAG | 2714205 | 1.7026331 | 5.2240915 | 39 |
| GGGGA | 1965185 | 1.7017909 | 15.441489 | 1 |
| GCCTC | 1803210 | 1.6837745 | 5.965769 | 15 |
| GACAG | 2679345 | 1.639218 | 6.226292 | 31 |
| GCAGT | 2659780 | 1.6277119 | 9.640218 | 22 |
| TTGCT | 3814325 | 1.609282 | 5.243939 | 41 |
| CGCCC | 1157165 | 1.6074761 | 6.3038244 | 25 |
| ACAGC | 2555695 | 1.6031989 | 5.0877466 | 14 |
| ACAGG | 2609610 | 1.5965542 | 6.23572 | 32 |
| AAATG | 5569005 | 1.5780073 | 5.2512116 | 42 |
| CTAAA | 5427755 | 1.5769647 | 5.0644903 | 40 |
| ACTTT | 5244675 | 1.5246422 | 5.2575336 | 33 |
| GAAGT | 3699745 | 1.521487 | 6.89717 | 46 |
| ATGAA | 5279710 | 1.4960341 | 5.245607 | 44 |
| AGGCC | 1642530 | 1.495398 | 7.152387 | 32 |
| CAGTT | 3542065 | 1.4939883 | 7.119596 | 23 |
| GGTGA | 2449945 | 1.4622372 | 6.1514 | 35 |
| CAGGT | 2374925 | 1.4533886 | 6.1030784 | 33 |
| ATCAA | 4993785 | 1.4508806 | 5.5637302 | 11 |
| GGGAC | 1621310 | 1.4395912 | 14.665741 | 2 |
| GTGCT | 2276195 | 1.3933659 | 5.291031 | 6 |
| GCCAT | 2218560 | 1.3921094 | 5.679182 | 2 |
| GCAAT | 3292790 | 1.3884519 | 6.9619093 | 12 |
| AGTTT | 4820700 | 1.3667501 | 6.267488 | 24 |
| AACTT | 4605595 | 1.3384783 | 5.12763 | 32 |
| AAGGC | 2171845 | 1.3287305 | 5.645146 | 31 |
| AAACT | 4554015 | 1.3231108 | 5.1925917 | 31 |
| GTTTA | 4659500 | 1.3210473 | 6.1187468 | 25 |
| GAGGT | 2165095 | 1.292226 | 5.4389467 | 3 |
| AATCA | 4411950 | 1.2818356 | 5.6707807 | 10 |
| AATGA | 4341630 | 1.2302241 | 5.111223 | 43 |
| GGACT | 1970355 | 1.205803 | 10.79102 | 3 |
| AAGCC | 1807285 | 1.1337181 | 5.4911537 | 46 |
| TCTAA | 3813380 | 1.1082448 | 5.073689 | 39 |
| CTAGC | 1708590 | 1.0721118 | 9.512642 | 9 |
| ATAGC | 2507680 | 1.0573992 | 7.1164026 | 19 |
| ACTAC | 2111665 | 0.9129819 | 7.121857 | 5 |
| CTACT | 2074245 | 0.8970591 | 6.865069 | 6 |
| GAACG | 1447690 | 0.88569385 | 5.3666735 | 38 |
| ACTAG | 1992865 | 0.84031993 | 6.4206476 | 8 |
| GACTA | 1498410 | 0.631826 | 6.8684177 | 4 |