![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | filtered_174gm_A_BiGosperm_L006_R1.fastq |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 171582441 |
| Filtered Sequences | 0 |
| Sequence length | 76 |
| %GC | 18 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
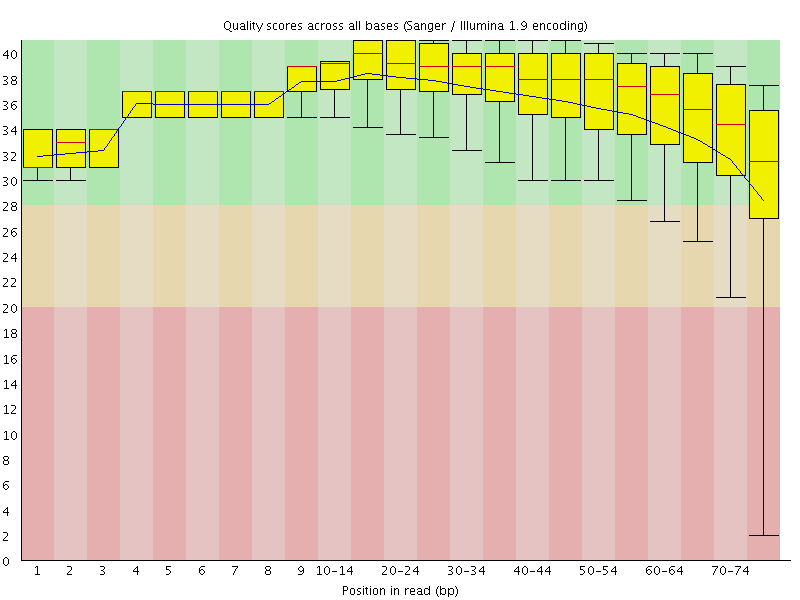
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
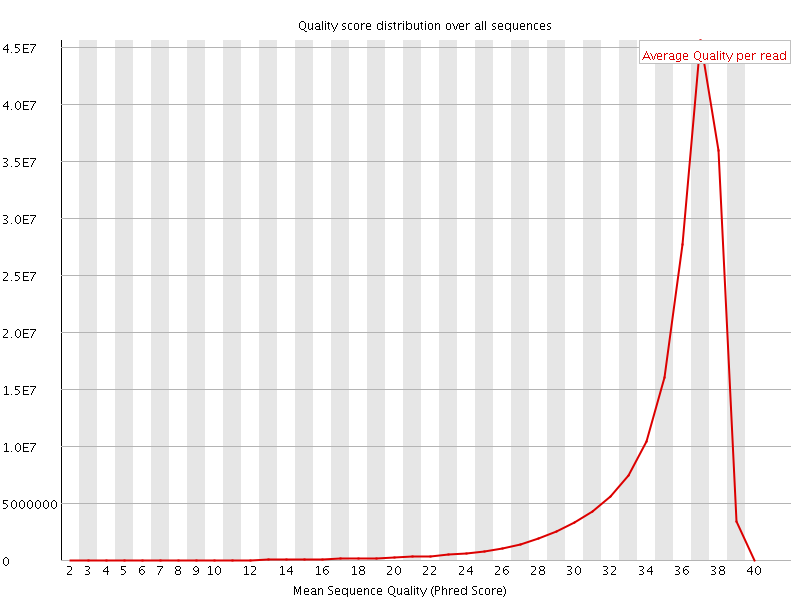
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
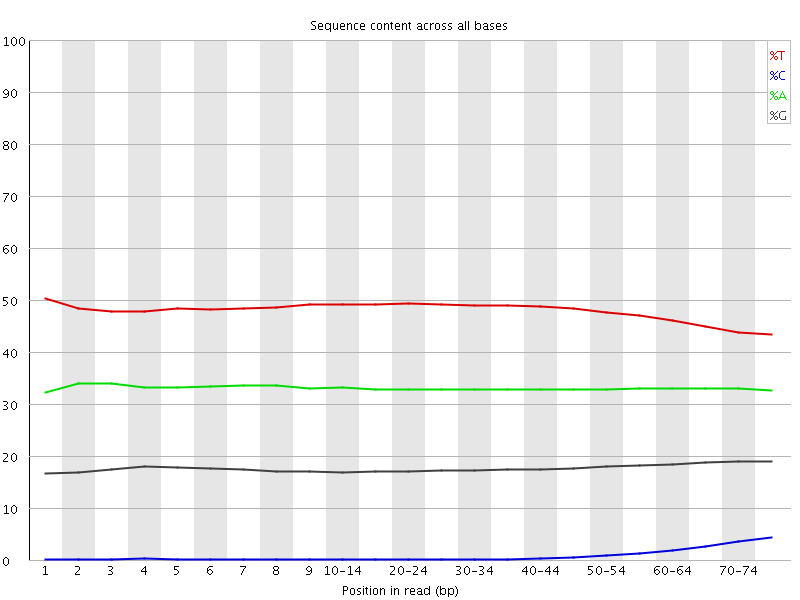
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
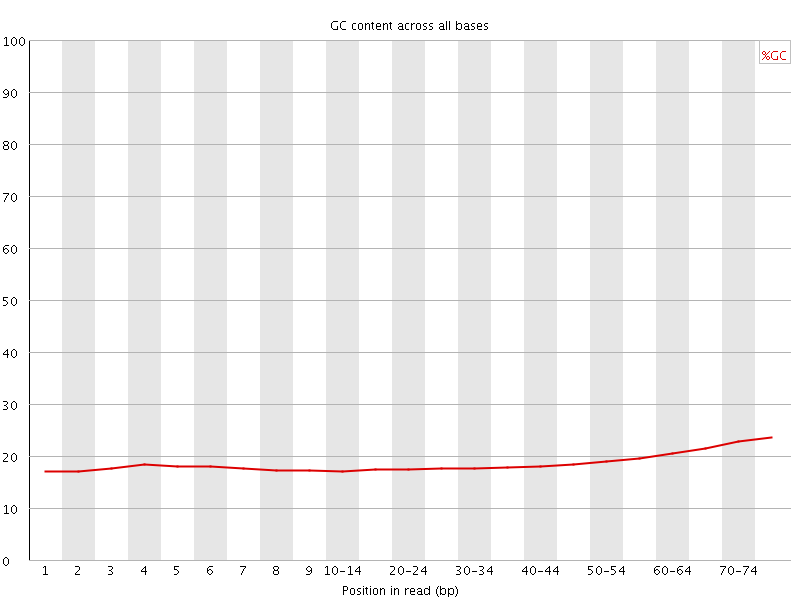
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
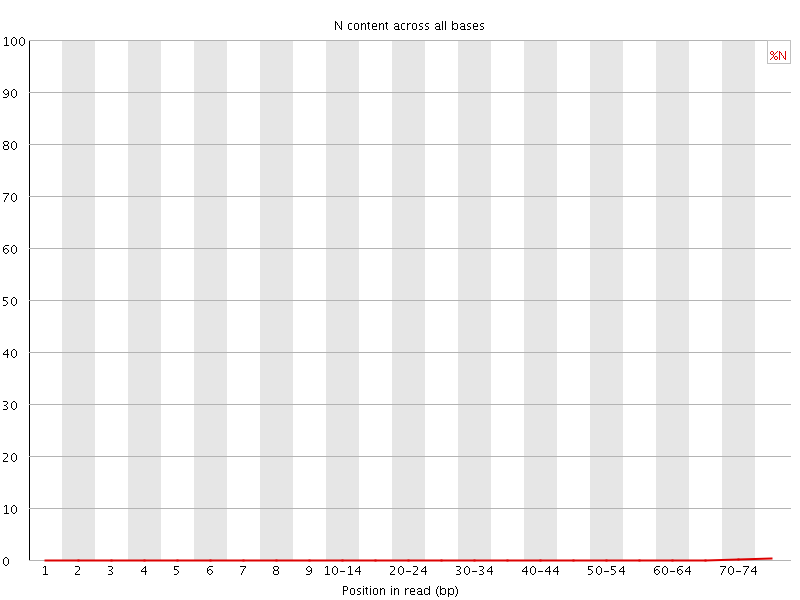
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
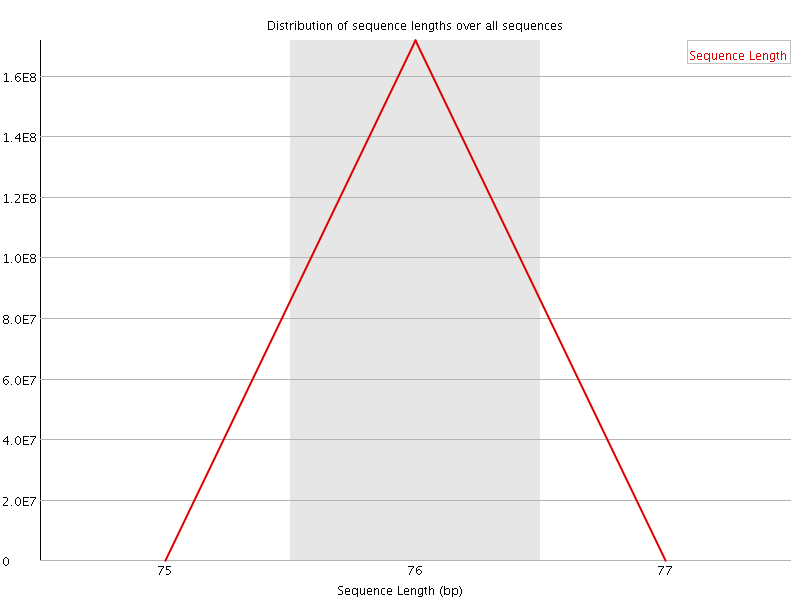
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
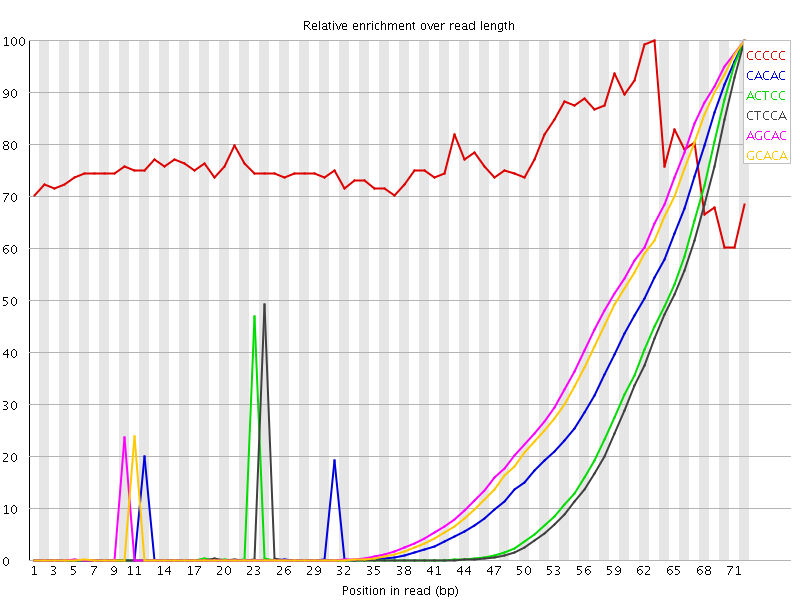
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCCCC | 40100 | 21939.352 | 28547.258 | 63 |
| CACAC | 13316205 | 7777.7886 | 43682.53 | 72 |
| ACTCC | 4301950 | 1734.989 | 12422.333 | 72 |
| CTCCA | 3862030 | 1557.568 | 11791.825 | 72 |
| AGCAC | 13804235 | 488.1336 | 2315.5806 | 72 |
| GCACA | 12943395 | 457.69327 | 2273.3809 | 72 |
| ACACG | 11005635 | 389.17188 | 2084.1794 | 72 |
| CACGT | 10216710 | 249.45554 | 1404.7568 | 72 |
| ACGTC | 9417450 | 229.94046 | 1310.0906 | 72 |
| CCCCG | 5705 | 188.96692 | 825.6892 | 71 |
| CCGTC | 242615 | 181.30159 | 11116.82 | 52 |
| CGTCT | 8834680 | 148.94594 | 856.44324 | 72 |
| GCCCC | 4075 | 134.97636 | 706.0809 | 72 |
| ACCCC | 7065 | 126.296036 | 729.7824 | 71 |
| CTCCC | 9330 | 115.16351 | 833.9638 | 72 |
| CACCC | 6325 | 113.06757 | 923.60425 | 72 |
| CCCGC | 2885 | 95.55996 | 382.9591 | 72 |
| TCCAG | 3560105 | 86.92503 | 681.68256 | 72 |
| CCCAC | 4540 | 81.15839 | 768.67584 | 70 |
| CCAGT | 3169775 | 77.39457 | 626.24664 | 72 |
| CCGCC | 2270 | 75.18929 | 250.29639 | 52 |
| CCCCA | 4110 | 73.47158 | 602.2543 | 69 |
| CACGC | 67030 | 72.54333 | 455.931 | 72 |
| CGCCC | 2090 | 69.22714 | 227.3637 | 71 |
| CAGTC | 2813475 | 68.695 | 556.9519 | 72 |
| CGCAC | 62895 | 68.06821 | 405.45657 | 71 |
| AACTC | 4959685 | 65.35572 | 459.19278 | 72 |
| CCACC | 3050 | 54.522694 | 568.32605 | 71 |
| GTCAC | 2230350 | 54.457176 | 456.7634 | 72 |
| CGCGC | 23830 | 47.786457 | 91.53153 | 69 |
| CGGAA | 21237070 | 45.464417 | 177.26395 | 72 |
| CCTCC | 3655 | 45.114967 | 352.31625 | 72 |
| GATCG | 25352225 | 37.475628 | 131.96066 | 72 |
| ATCGG | 24454490 | 36.148598 | 129.62354 | 72 |
| AGAGC | 16268745 | 34.828205 | 153.56142 | 72 |
| TCGGA | 23039310 | 34.05668 | 128.91801 | 72 |
| CACTG | 1334420 | 32.581764 | 361.30966 | 33 |
| GAGCA | 15157820 | 32.449932 | 150.33292 | 72 |
| TCACA | 1972045 | 25.98641 | 225.06862 | 72 |
| GCACC | 20515 | 22.20239 | 143.11383 | 72 |
| ACACT | 1545945 | 20.371523 | 194.71552 | 32 |
| AGATC | 25529100 | 20.366468 | 70.210495 | 71 |
| CTCCG | 27075 | 20.232635 | 198.44661 | 72 |
| CGGCG | 157945 | 19.175089 | 23.240093 | 24 |
| CGCGG | 154845 | 18.798738 | 22.26944 | 4 |
| CCCCT | 1500 | 18.515034 | 205.14618 | 72 |
| TCGCG | 380875 | 17.231266 | 20.177341 | 3 |
| CCACG | 15570 | 16.850657 | 94.61957 | 71 |
| CGTCG | 370915 | 16.780663 | 21.280561 | 71 |
| GCCAC | 15490 | 16.764076 | 105.57577 | 72 |
| GCTCC | 22145 | 16.548538 | 130.66724 | 71 |
| CACCG | 14545 | 15.741349 | 89.536705 | 71 |
| CGCGA | 237545 | 15.564128 | 18.051582 | 4 |
| ACGCG | 229995 | 15.069447 | 28.197424 | 70 |
| CCGGC | 7350 | 14.739004 | 355.74014 | 52 |
| GCGCG | 119775 | 14.541114 | 17.371824 | 70 |
| CGCGT | 313275 | 14.172955 | 22.669844 | 71 |
| TCCCC | 1100 | 13.577692 | 98.11339 | 72 |
| CGACG | 206540 | 13.532658 | 16.584127 | 1 |
| GGCCC | 6555 | 13.144787 | 63.753036 | 71 |
| CCCAG | 11160 | 12.077928 | 79.761955 | 71 |
| CCCGG | 5515 | 11.059267 | 60.85517 | 71 |
| TGCCG | 237970 | 10.766063 | 684.3324 | 50 |
| GCCGT | 235075 | 10.635089 | 680.54535 | 51 |
| CGTCC | 14005 | 10.465671 | 59.945377 | 70 |
| ACGCC | 7965 | 8.620134 | 60.212456 | 71 |
| GTCTG | 8181415 | 8.350587 | 50.136734 | 72 |
| GGCAC | 120155 | 7.872647 | 43.46342 | 72 |
| CCCTC | 630 | 7.7763143 | 57.976093 | 72 |
| GCACG | 113940 | 7.465435 | 42.702454 | 71 |
| ACACC | 12405 | 7.2455664 | 40.725864 | 71 |
| CGGAC | 103605 | 6.788278 | 8.365942 | 4 |
| ACCCG | 6215 | 6.726194 | 44.96385 | 71 |
| CGCTC | 8925 | 6.669483 | 393.2453 | 2 |
| CGCCG | 3300 | 6.6175127 | 54.83971 | 50 |
| AGCGC | 98975 | 6.484918 | 26.53731 | 72 |
| CACCA | 11055 | 6.4570537 | 34.39819 | 72 |
| ACCAC | 11015 | 6.4336905 | 36.72346 | 70 |
| GGGGG | 13331345 | 5.9320793 | 6.657552 | 43 |
| CGGCC | 2955 | 5.9256816 | 34.7809 | 70 |
| CTCAC | 14560 | 5.872091 | 46.487995 | 70 |
| ATGCC | 239535 | 5.8485885 | 366.9148 | 49 |
| TCTCG | 337120 | 5.683585 | 254.03279 | 43 |
| GCGCC | 2825 | 5.664992 | 23.18479 | 72 |
| CCAGC | 5205 | 5.633119 | 47.704605 | 72 |
| CCACA | 9330 | 5.449507 | 31.863241 | 71 |
| CACAT | 412355 | 5.433764 | 35.75541 | 72 |
| CTGAA | 6798260 | 5.4234796 | 33.904675 | 72 |
| CACTC | 13420 | 5.4123254 | 36.37895 | 69 |
| GCGTC | 118940 | 5.3809958 | 13.207432 | 72 |
| CCGCG | 2660 | 5.3341165 | 15.213793 | 71 |
| CTCGT | 314980 | 5.310322 | 254.54828 | 44 |
| TGAAC | 6403715 | 5.108721 | 31.366726 | 72 |
| GACCC | 4700 | 5.086583 | 40.275208 | 72 |
| GCCCG | 2410 | 4.8327894 | 18.836124 | 71 |
| GCCCA | 4370 | 4.7294393 | 29.326605 | 72 |
| AACCC | 8045 | 4.6989594 | 36.297474 | 72 |
| CCCGT | 6245 | 4.6667705 | 39.68932 | 72 |
| CGCCA | 4305 | 4.659093 | 33.23682 | 72 |
| GCGGC | 38065 | 4.621228 | 6.2907033 | 52 |
| CGGTC | 100400 | 4.5422225 | 10.395113 | 71 |
| GCCGC | 2260 | 4.5319934 | 16.596373 | 51 |
| CGGGC | 37250 | 4.522284 | 6.025849 | 4 |
| GAACT | 5667620 | 4.5214834 | 30.590212 | 72 |
| CATAC | 335390 | 4.4195657 | 25.619156 | 72 |
| GCGCA | 65035 | 4.2611423 | 22.03946 | 72 |
| TCTGA | 7473960 | 4.1170554 | 25.248346 | 72 |
| ACGGC | 61110 | 4.0039735 | 10.771164 | 72 |
| TCGGC | 88435 | 4.000911 | 6.593453 | 58 |
| TCTGC | 230445 | 3.8851264 | 251.40291 | 58 |
| CTGCT | 229060 | 3.8617764 | 251.28369 | 59 |
| CGGGG | 522530 | 3.8405504 | 4.654858 | 1 |
| GGAAG | 28730035 | 3.7236078 | 11.735041 | 72 |
| TTCGC | 211995 | 3.5740736 | 4.2873855 | 2 |
| ATCTC | 382705 | 3.4821656 | 136.56407 | 42 |
| CGTTC | 206335 | 3.4786503 | 4.111262 | 4 |
| GTCGC | 75605 | 3.4204657 | 7.110437 | 72 |
| TCCCA | 8245 | 3.3252327 | 26.228666 | 72 |
| GGCGC | 27155 | 3.2967148 | 6.711125 | 72 |
| TCGTC | 195240 | 3.2915971 | 3.9254644 | 55 |
| CGAGC | 48695 | 3.1905332 | 3.8827689 | 70 |
| AGCCC | 2920 | 3.1601748 | 19.94049 | 71 |
| TACAC | 232525 | 3.0640733 | 20.063023 | 72 |
| CTTCC | 10000 | 2.784756 | 145.13197 | 6 |
| CGATC | 108165 | 2.6410027 | 15.605688 | 10 |
| CGCCT | 3445 | 2.574383 | 20.249653 | 72 |
| CGGAG | 645255 | 2.5595348 | 6.0418177 | 71 |
| CAGCC | 2365 | 2.559525 | 16.815718 | 70 |
| CACGG | 37940 | 2.485858 | 16.617031 | 71 |
| GAAGA | 34810900 | 2.4349523 | 6.7620263 | 72 |
| TCTCC | 8700 | 2.4227376 | 21.93214 | 71 |
| CACAG | 64795 | 2.2912257 | 22.920343 | 72 |
| CTCCT | 7795 | 2.1707172 | 39.94385 | 72 |
| GACAC | 59770 | 2.113536 | 13.133842 | 72 |
| AAGAG | 30135635 | 2.1079268 | 6.1319585 | 72 |
| ACCTC | 5210 | 2.1012082 | 16.173044 | 71 |
| CCGAC | 1915 | 2.0725117 | 40.111656 | 52 |
| AGTCA | 2595305 | 2.0704687 | 17.096888 | 72 |
| ACCGC | 1685 | 1.8235941 | 10.166557 | 72 |
| TCCCG | 2350 | 1.7561105 | 14.03976 | 72 |
| CACCT | 4005 | 1.6152282 | 14.425766 | 72 |
| CTTCT | 243255 | 1.5282795 | 94.333855 | 56 |
| TCTTC | 237100 | 1.48961 | 93.304665 | 55 |
| ACCCA | 2520 | 1.4718928 | 13.717069 | 72 |
| TCGTA | 2643455 | 1.4561558 | 9.7288065 | 45 |
| ACTCT | 158535 | 1.442482 | 14.30693 | 72 |
| ACTTC | 158030 | 1.4378871 | 10.536239 | 72 |
| GGGCC | 11745 | 1.4258853 | 7.1943927 | 70 |
| CAGAC | 39650 | 1.4020694 | 9.302003 | 70 |
| CAACC | 2370 | 1.38428 | 7.8288283 | 69 |
| CCCTG | 1820 | 1.3600516 | 12.148814 | 71 |
| TCACC | 3340 | 1.3470318 | 11.948614 | 72 |
| CGTAT | 2335460 | 1.2864959 | 9.511429 | 46 |
| CGGCT | 27460 | 1.2423251 | 9.881937 | 53 |
| CCCGA | 1140 | 1.2337668 | 7.820428 | 72 |
| CTCTA | 133525 | 1.2149204 | 9.822865 | 72 |
| TCCGG | 26040 | 1.1780825 | 10.657482 | 72 |
| ATTCC | 128840 | 1.1722926 | 8.165997 | 72 |
| CTCGC | 1560 | 1.1657584 | 5.684898 | 69 |
| CTTCA | 127335 | 1.1585987 | 8.649249 | 72 |
| AGCTC | 46870 | 1.1443979 | 7.1450377 | 71 |
| CCGGT | 25140 | 1.1373652 | 10.2815275 | 72 |
| CCACT | 2750 | 1.1090829 | 11.220039 | 72 |
| GAGCC | 16250 | 1.0647125 | 5.1375666 | 70 |
| TTCCA | 115760 | 1.0532799 | 7.975324 | 72 |
| GTCCC | 1365 | 1.0200386 | 8.640775 | 70 |
| CGACC | 930 | 1.006494 | 5.08287 | 71 |
| TCCAC | 2485 | 1.0022078 | 9.471462 | 72 |
| ACTCG | 40785 | 0.99582386 | 7.5596256 | 71 |
| CCGCT | 1330 | 0.9938837 | 5.4004846 | 70 |
| TGCAC | 39455 | 0.96335006 | 6.4303694 | 69 |
| ACTGA | 1191625 | 0.9506481 | 12.053662 | 34 |
| CCAAC | 1540 | 0.89948994 | 6.54146 | 71 |
| CAGGC | 13340 | 0.8740469 | 5.6104746 | 72 |
| GCCGG | 7050 | 0.8558953 | 22.192171 | 51 |
| CCAGG | 12155 | 0.7964048 | 5.77572 | 71 |
| ATCCC | 1855 | 0.748127 | 8.596483 | 71 |
| CTCGA | 30065 | 0.73407984 | 7.357334 | 72 |
| ACCCT | 1800 | 0.7259453 | 8.597174 | 72 |
| GACTC | 28125 | 0.68671197 | 5.2401156 | 72 |
| GGCCG | 5650 | 0.68593025 | 11.314412 | 50 |
| CTGAT | 1062000 | 0.58500624 | 8.332965 | 35 |
| GCACT | 23610 | 0.5764718 | 7.1544333 | 72 |
| ACGCT | 21875 | 0.5341093 | 13.081073 | 1 |
| GCTCG | 8610 | 0.38952726 | 5.8930535 | 43 |
| ACTCA | 29515 | 0.38893074 | 7.898565 | 72 |
| AGGCC | 5730 | 0.375434 | 5.8469915 | 49 |
| TATGC | 536600 | 0.29558787 | 8.491481 | 48 |
| TCCGA | 10905 | 0.26626113 | 12.962193 | 8 |
| GCTTG | 247085 | 0.2521941 | 15.427195 | 61 |
| CCGAT | 9770 | 0.23854846 | 12.944729 | 9 |
| TTCCG | 12785 | 0.21554531 | 8.980506 | 7 |
| GCTCT | 11950 | 0.20146786 | 9.047201 | 3 |
| CCGTG | 3850 | 0.17417884 | 5.3559403 | 52 |
| CTTGA | 235060 | 0.12948357 | 7.669823 | 62 |
| TTCTG | 274825 | 0.10453168 | 5.7800326 | 57 |
| GTCTT | 274265 | 0.104318686 | 5.7426353 | 54 |
| TGCTT | 265870 | 0.10112558 | 5.7383904 | 60 |