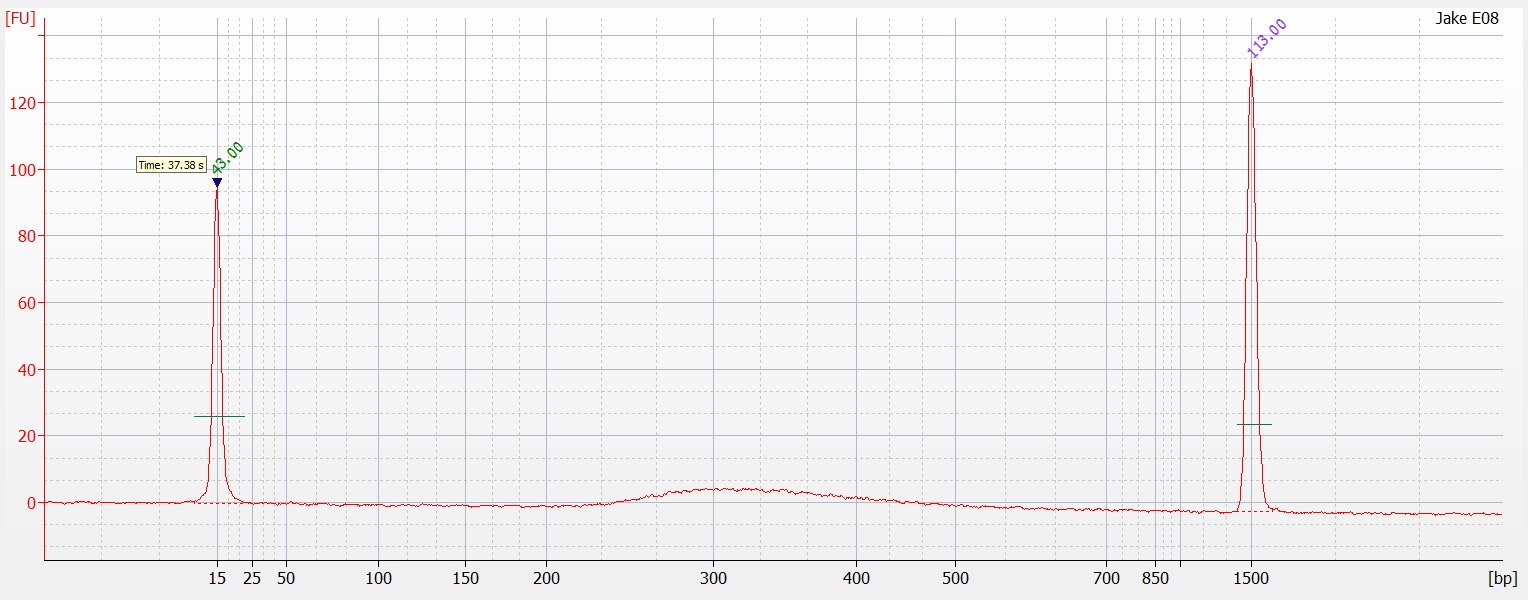
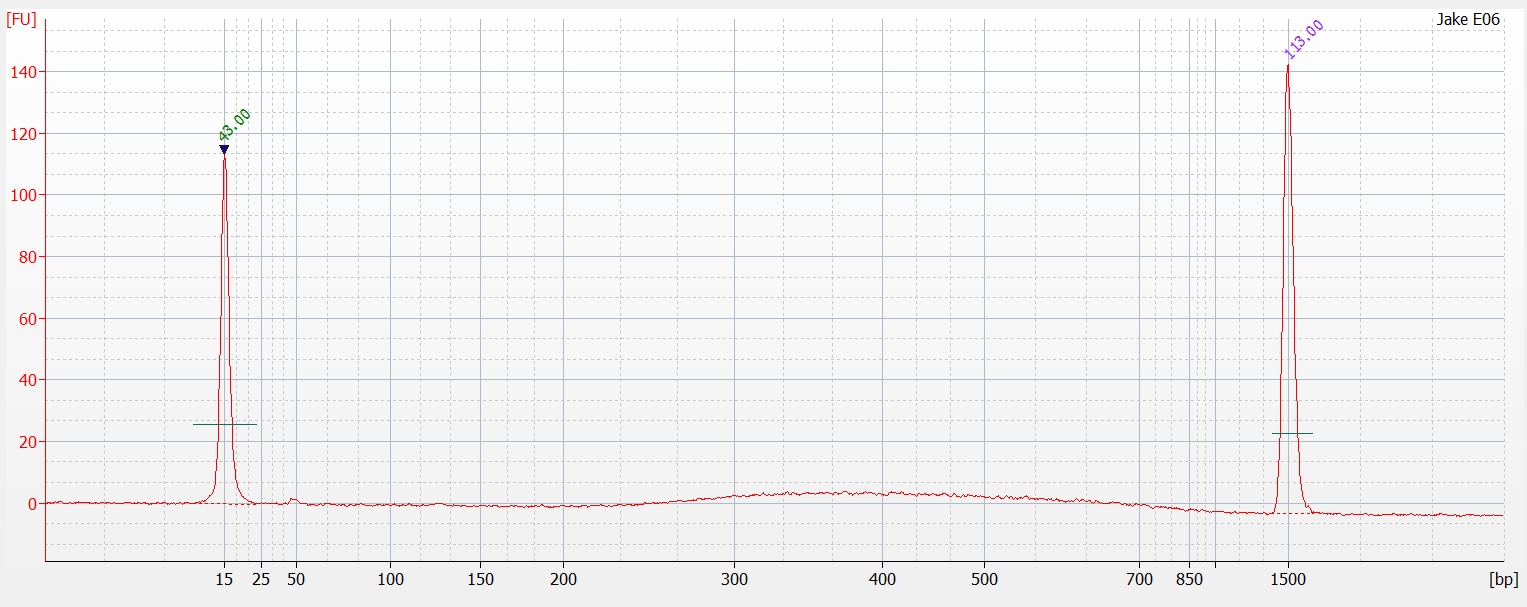
Today I ran half of the 96 well plate extraction (
part 1 and
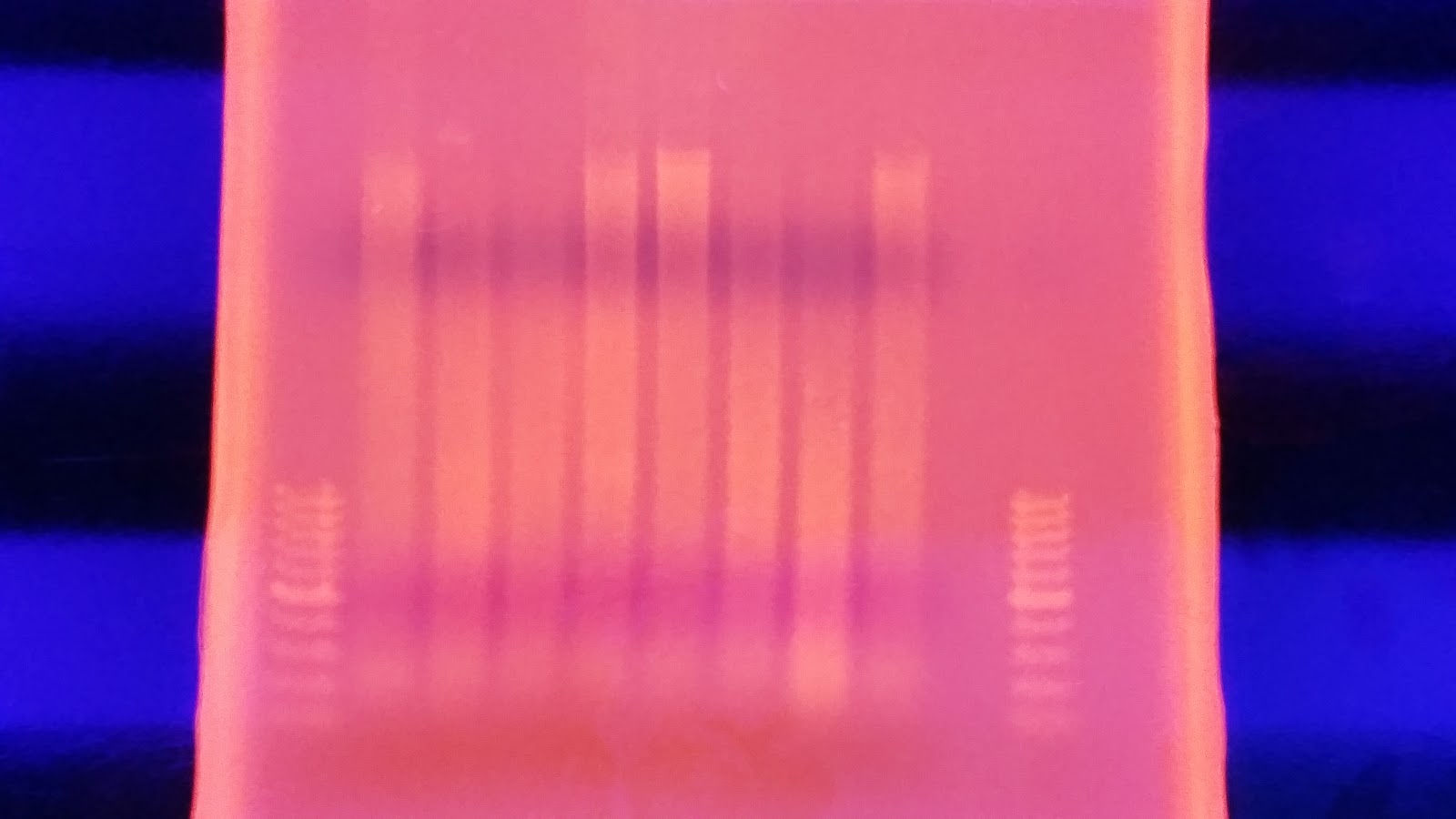
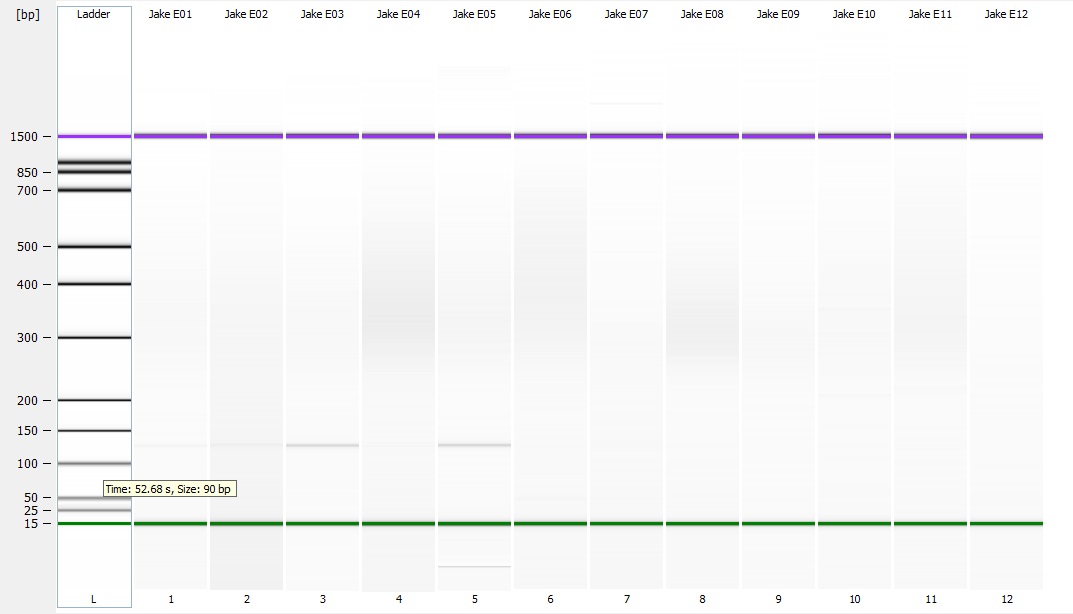
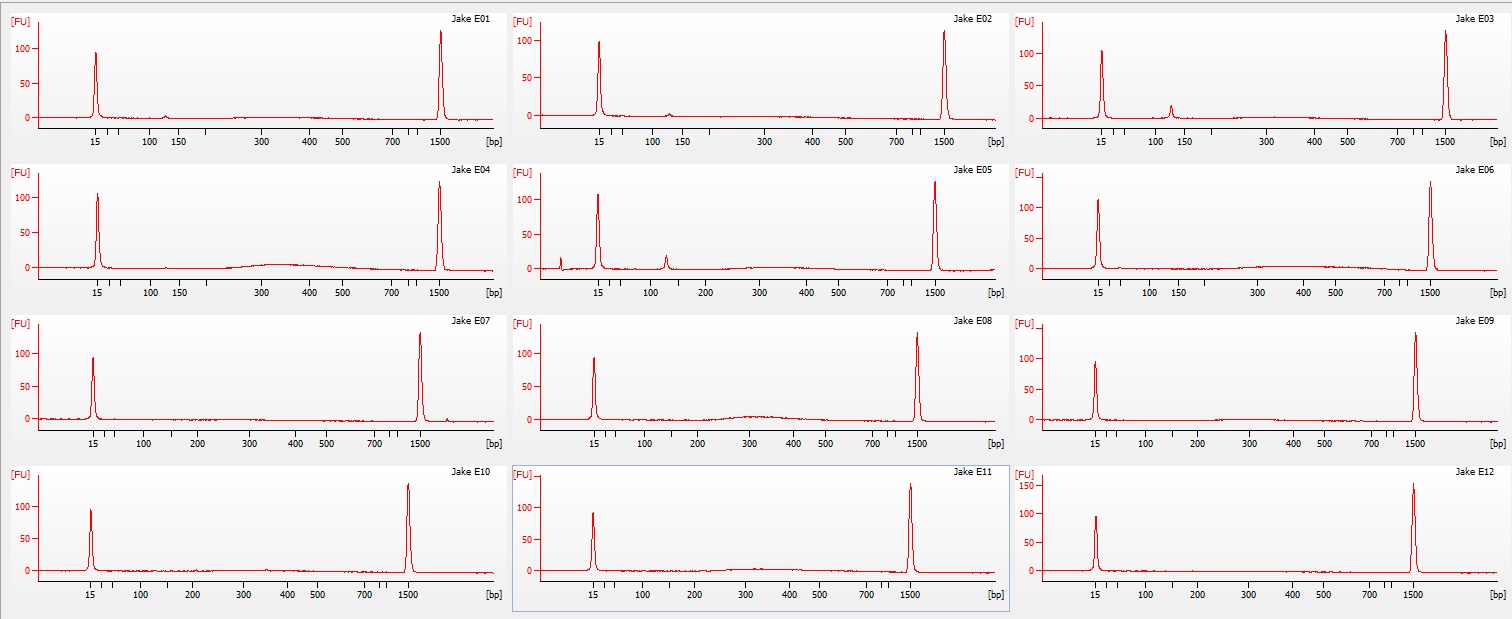
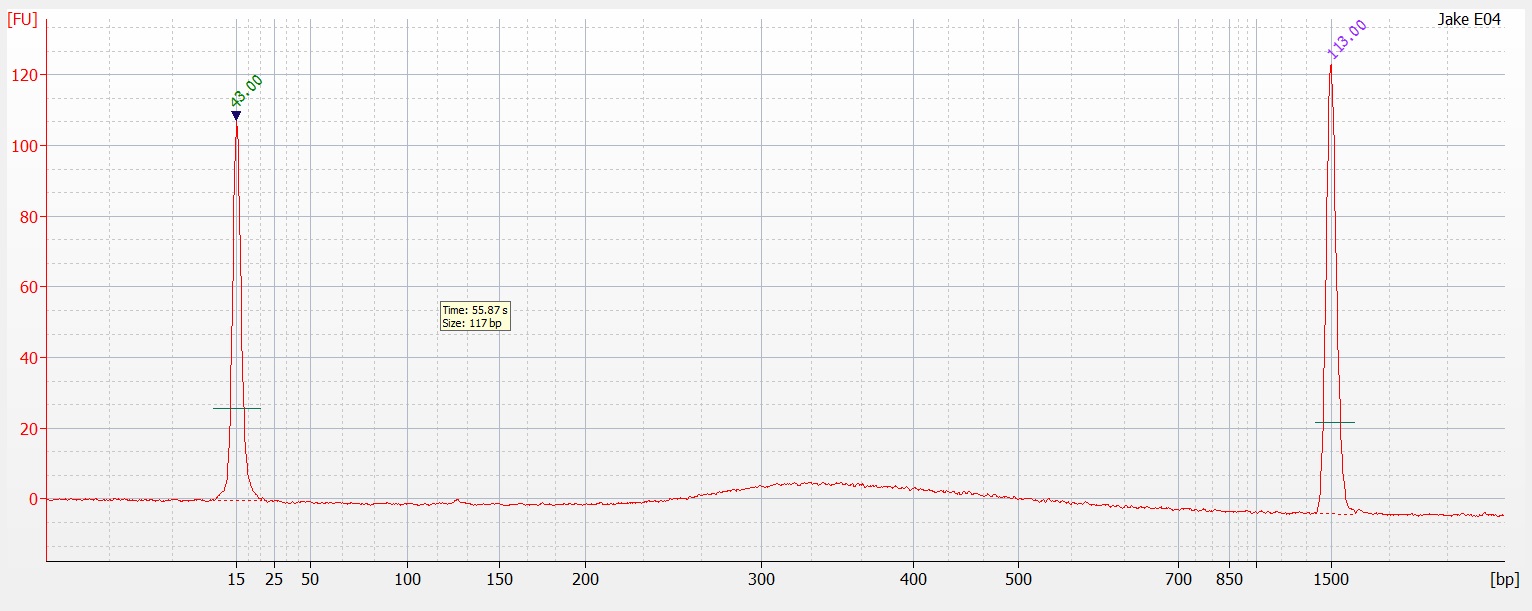
part 2) on a 1.3% agarose gel to determine the quality of the DNA isolated. Sadly the DNA is heavily degraded and there appears to be no high molecular weight band. To fit all of the samples, I did not add a ladder which would tell us exactly what the weight of the DNA is. Overall this is very disappointing. I'm going to run the other half tomorrow to see if maybe they are of better quality or may there was a loading issue with the samples.
Protocol for today:
Used a super sized gel mold and wells. So it took a pretty big volume.
Gel:
500 ml 1X Low TAE
6.5 g general purpose agarose
Microwave solution for 3 minutes at a time until boiling.
Visually inspect for dense materials (apparently I didn't do well enough as one corner of the gel was higher density than the others.
Placed two rows of 24 well combs in gel mold.
Slowly poured gel into one corner of the mold.
Allowed get to set for 30 minutes.
Once gel was firm I placed it in the electrophoresis chamber.
I filled the chamber with 1 X Low TAE solution until it was even with the gels top surface.
Pipetted 30 ul of Low TAE into each well. Then loaded wells with DNA using the 12 channel pipette in a left to right, top to bottom pattern.
Ran gel for 10 minutes at 120 v.
Paused gel and added 1 ul 5X loading dye to the left most well on both rows.
Ran gel for another 10 minutes at 120 v.
Paused gel and added 1X Low TAE until gel was thoroughly covered.
Ran gel for 1.5 hours until dye had moved appropriately far down the lane.
Placed gel in EtBr bath made 1 L water and 1 ml EtBr.
Soaked gel for 30 minutes.
Imaged on the translinker.
Samples organized:
| Well | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | 18 | 19 | 20 | 21 | 22 | 23 | 24 |
| Top Row | 1N1-4 1 | 1N1-4 2 | 1N1-4 3 | 1N1-4 4 | 1N1-4 5 | 1N1-4 6 | 1N1-4 7 | 1N1-4 8 | 1N1-4 11 | 1N1-4 10 | 1H1-4 1 | 1H1-4 2 | 1H1-4 3 | 1H1-4 4 | 1H1-4 5 | 1H1-4 6 | 1H1-4 7 | 1H1-4 8 | 1H1-4 9 | 1H1-4 10 | 1S1-4 1 | 1S1-4 2 | 1S1-4 3 | 1S1-4 4 |
| Bottom Row | 1S1-4 5 | 1S1-4 6 | 1S1-4 7 | 1S1-4 8 | 1S1-4 9 | 1S1-4 10 | 2N1-4 1 | 2N1-4 2 | 2N1-4 3 | 2N1-4 4 | 2N1-4 5 | 2N1-4 6 | 2N1-4 7 | 2N1-4 8 | 2N1-4 9 | 2N1-4 10 | 2H1-4 1 | 2H1-4 2 | 2H1-4 3 | 2H1-4 4 | 2H1-4 5 | 2H1-4 6 | 2H1-4 7 | 2H1-4 8 |
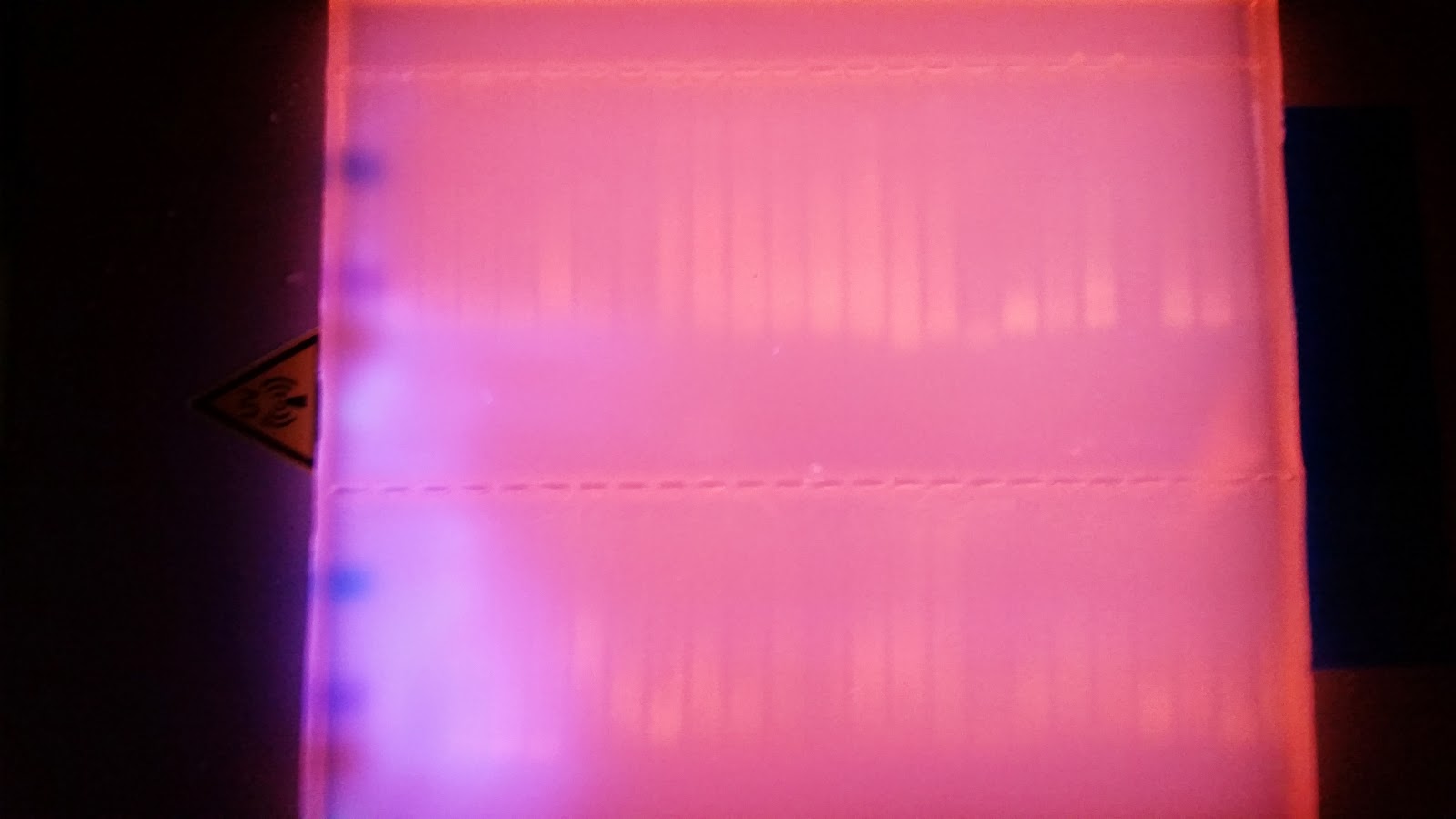 |
| Full Gel |
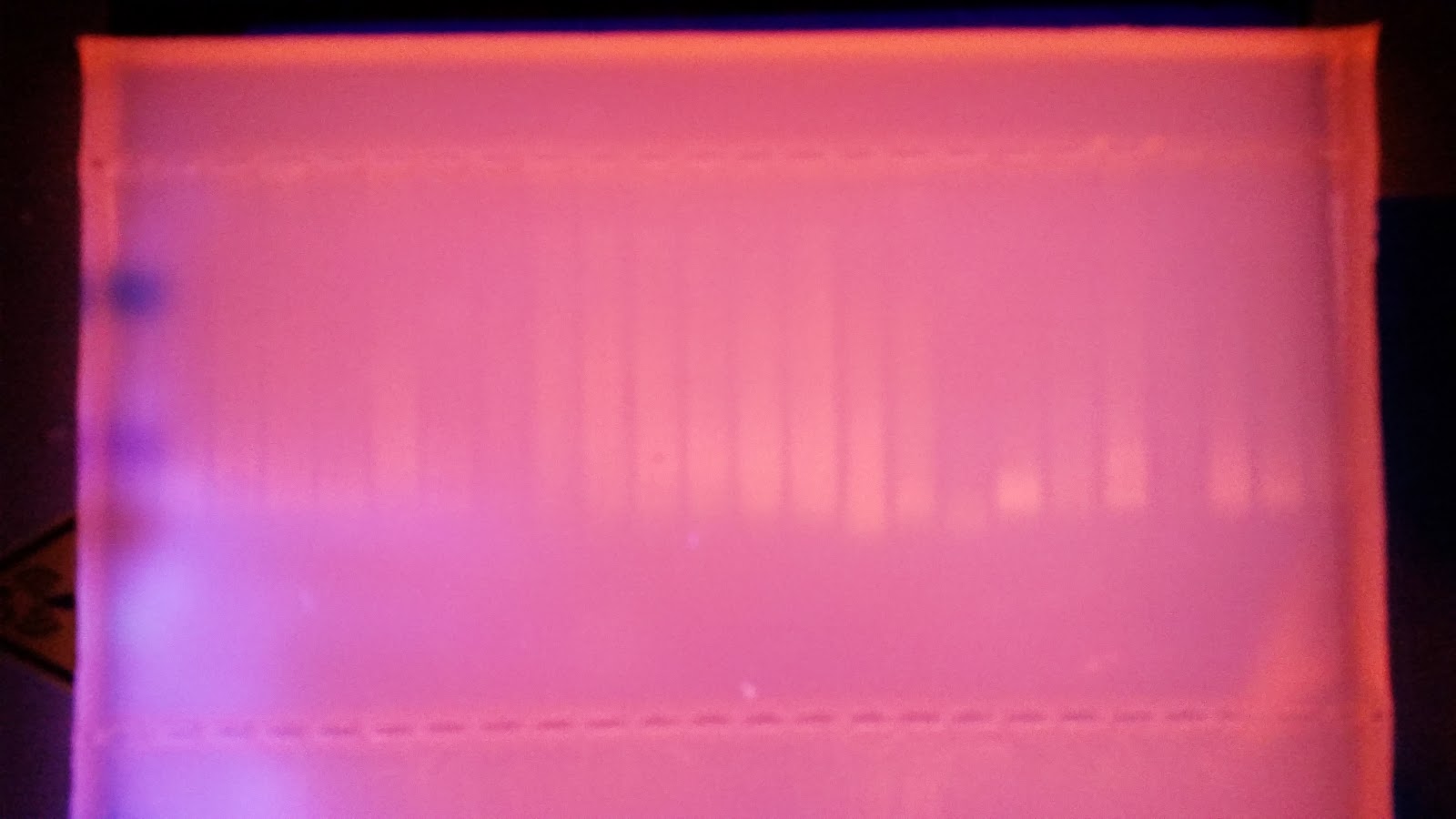 |
| Top Row Samples A1-12, B1-12 |
 |
| Bottom Row Samples C1-12, D1-12 |