Tanner crab RNA has proved a bit troublesome. As such, Steven asked me to try isolating some RNA using the RNeasy Plus Mini Kit (Qiagen) to see how things would turn out.
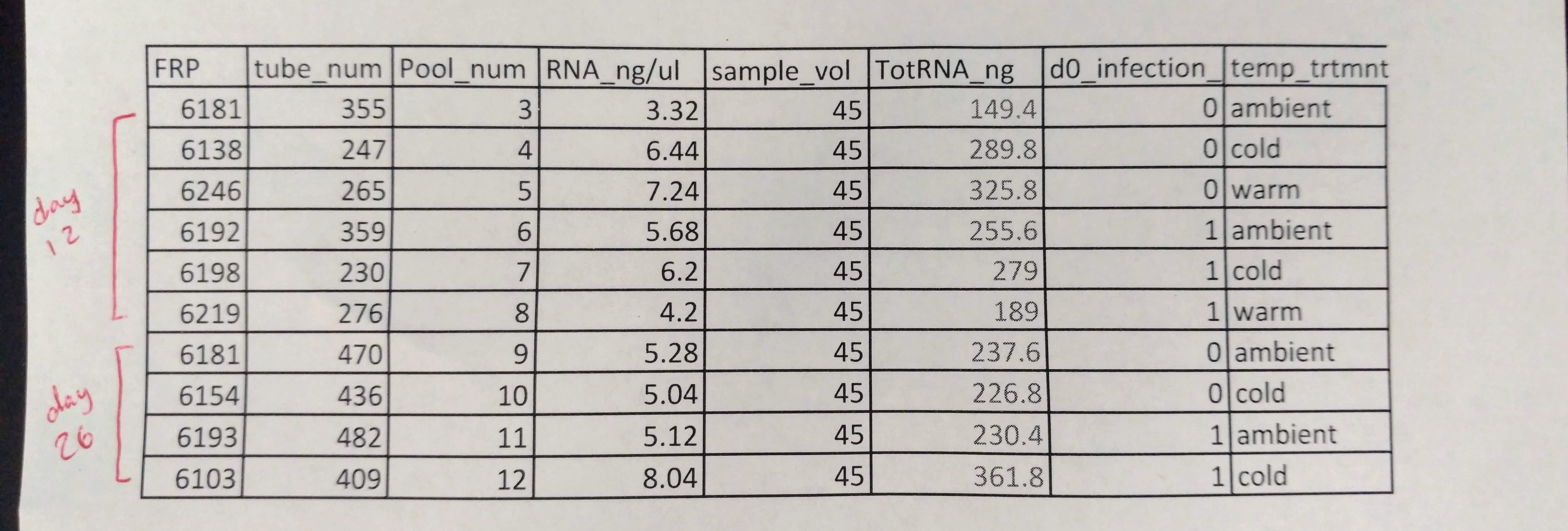
Grace provided me with the following samples:

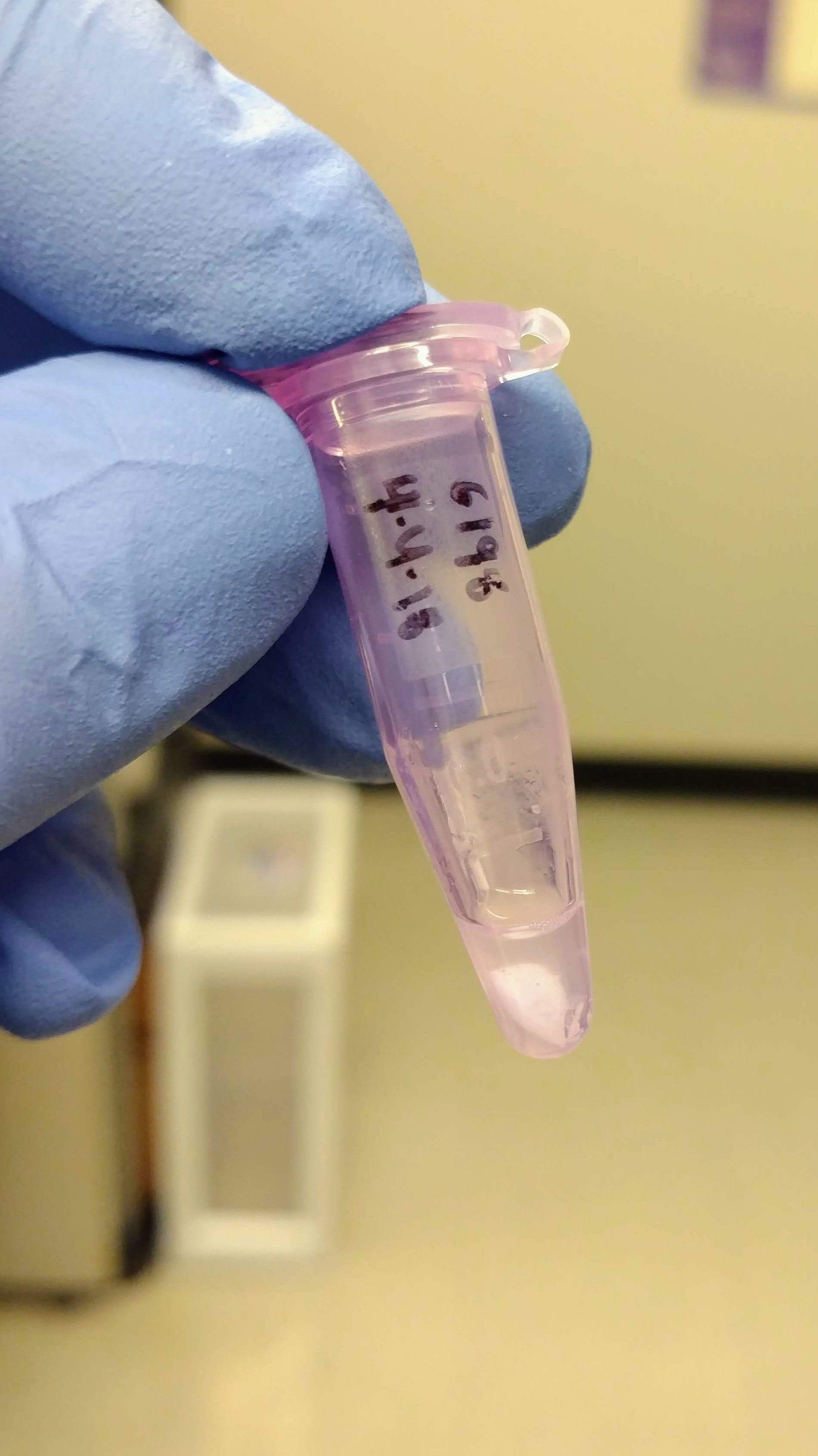
Crab hemolymph had been collected (100uL?) and preserved with 1mL (?) of RNAlater. Grace pelleted the samples, removed the supernatant, and stored the pelleted material at -80C. Here’s what that looked like:
RNA was isolated according to the manufacturer’s protocol – following guideline for samples with < 1 x 106 cells.
One interesting thing that happened is a precipitate formed after adding the initial buffer to the sample:
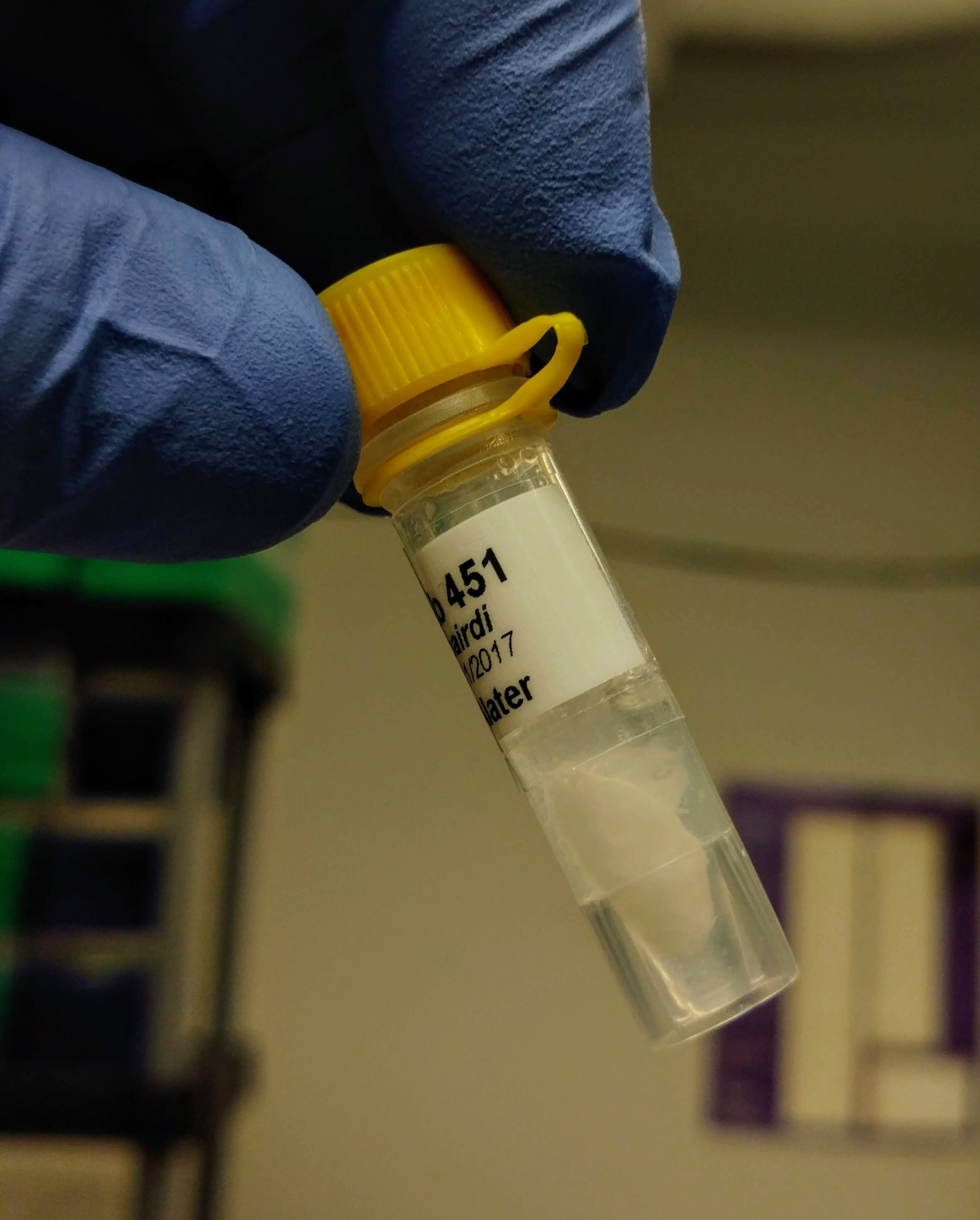
A solid precipitate formed in each of the tubes that could not be dispersed – it actually looked like a small piece of paper was now present in each tube.
Samples were spun and the supernatant was utilized (this was the normal progression of the protocol, regardless of this precipitate forming).
Samples were eluted with 30uL of nuclease-free water.
Samples were quantified using the Roberts Lab Qubit 3.0 with the RNA High Sensitivity asssay (Invitrogen). Used 5uL of sample for measurements.
Samples were also assessed with the Roberts Lab NandoDrop1000. Samples were recovered from the pedestal after measurement.
RNA was given to Grace for storage at -80C.
RESULTS
Qubit measurements (Google Sheet):
– 20180731_qubit_RNA_crab_isos
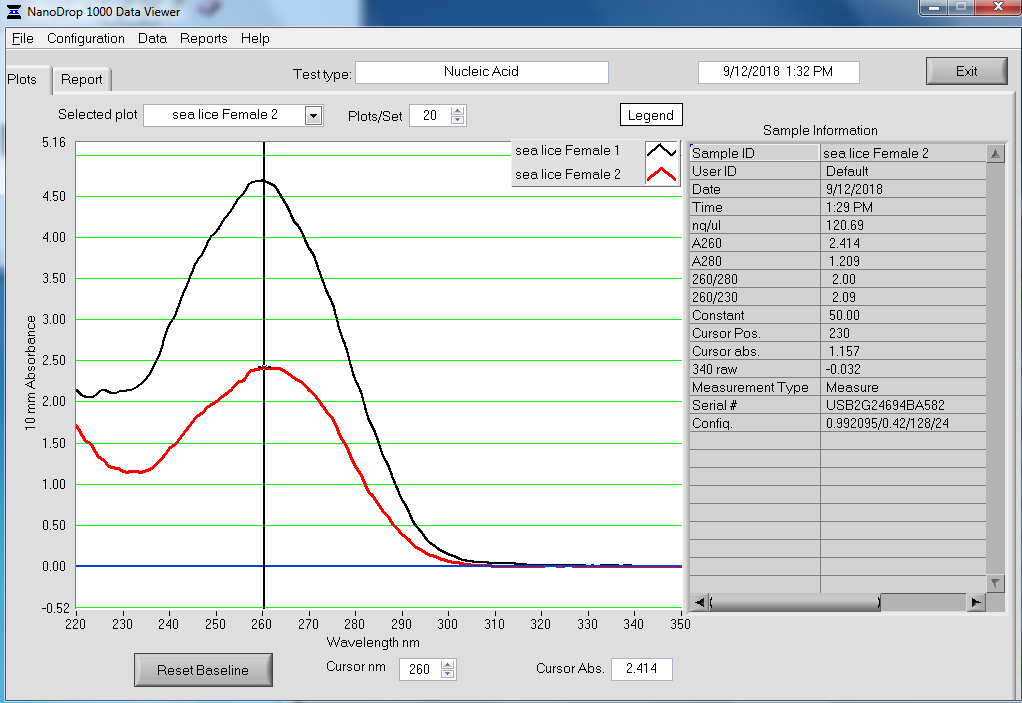
NanoDrop Spec Curves:
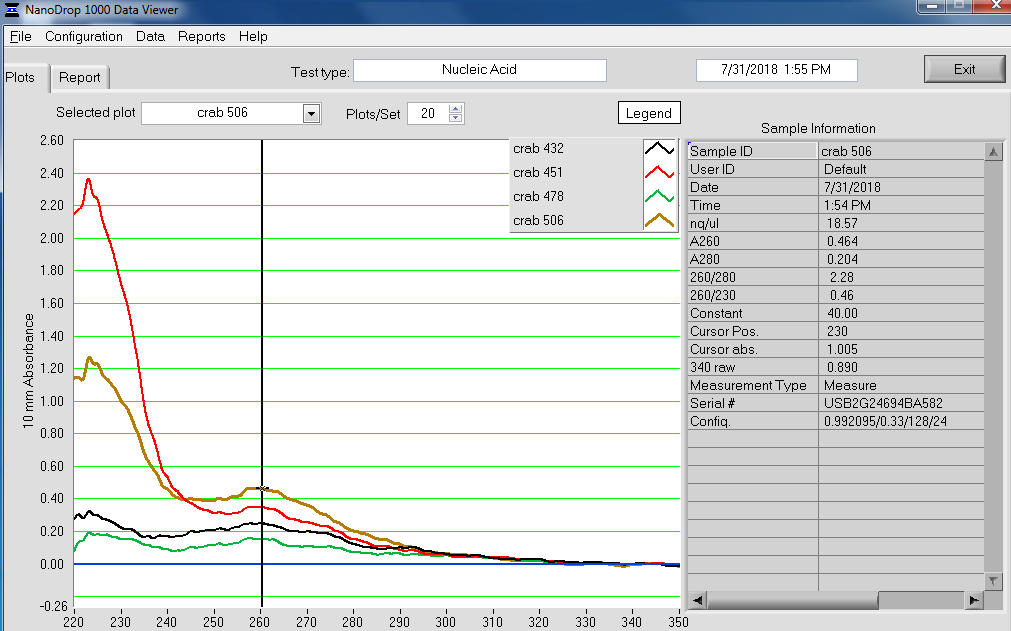
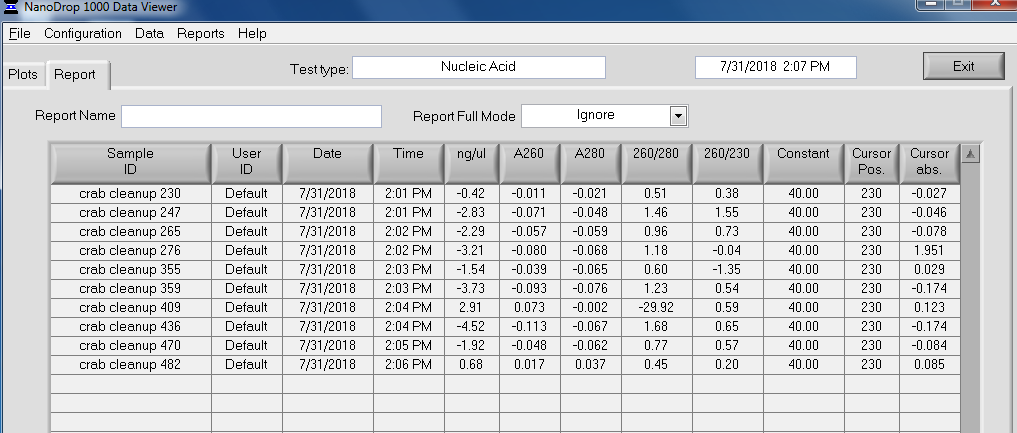
NanoDrop Table:
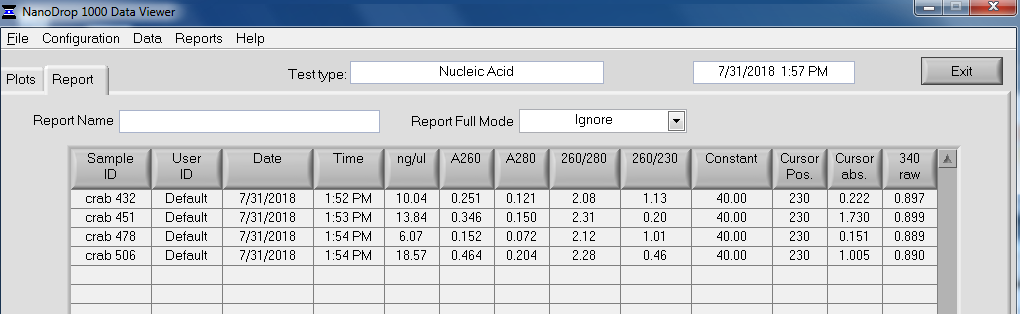
Overall, the isolation looks pretty good. The purity looks good (NanoDrop 260/280 ratios) and the absorbance peak at 260nm is exactly where we would want/expect it to be.
The yields (according to the Qubit) are OK. They range from ~37ng – 350ng.
The important part is that this method produced clean RNA, which means the quantification is believable. I think Grace’s earlier RNA isolations using RNAzol RT had too much contamination carried over, leading to incorrect quantification measurements.
Going forward, I think we need to use some sort of isolation kit, however, we will be testing out good, old TriReagent as well.