Redundans should assemble both Illumina and PacBio data, so let’s do that.
Sean had previously performed this – twice actually:
It wasn’t entirely clear how he had run Redundans the first time and the second time he used his Platinus contig FASTA file as the necessary reference assembly when running Redundans.
Since he had produced a good looking assembly from PacBio data using Canu, I decided to give Redundans a rip using that assembly.
I then compared all three Redundans runs using QUAST.
Jupyter notebook (GitHub): 20171004_docker_oly_redundans.ipynb
Notebook is also embedded at the bottom of this notebook entry (but, it should be easier to view at the link provided above).
- Sean’s Canu assembly (FASTA): http://owl.fish.washington.edu/scaphapoda/Sean/Oly_Canu_Output/oly_pacbio_.contigs.fasta
- Sean’s first Redundans assembly (scaffolded assembly; FASTA): http://owl.fish.washington.edu/scaphapoda/Sean/Oly_Redundans_Output/scaffolds.reduced.fa
- Sean’s second Redundans assembly (scaffolded assembly; FASTA): http://owl.fish.washington.edu/scaphapoda/Sean/Oly_Redundans_Output_Try_2/scaffolds.reduced.fa
-
Redundans Output folder: http://owl.fish.washington.edu/Athaliana/20171004_redundans/
- Redundans assembly (scaffolded assembly; FASTA): http://owl.fish.washington.edu/Athaliana/20171004_redundans/scaffolds.reduced.fa
- Quast Output folder (default settings): http://owl.fish.washington.edu/Athaliana/quast_results/results_2017_10_05_14_21_50/
- Quast Output folder (–scaffolds option): http://owl.fish.washington.edu/Athaliana/quast_results/results_2017_10_05_14_28_51/
Of note, is that Redundans didn’t find any alignments for the paired reads for each of the BGI mate-pair Illumina data:
- 160103_I137_FCH3V5YBBXX_L3_WHOSTibkDCABDLAAPEI-62_2.fq.gz
- 160103_I137_FCH3V5YBBXX_L3_WHOSTibkDCACDTAAPEI-75_2.fq.gz
- 160103_I137_FCH3V5YBBXX_L4_WHOSTibkDCABDLAAPEI-62_2.fq.gz
- 160103_I137_FCH3V5YBBXX_L4_WHOSTibkDCACDTAAPEI-75_2.fq.gz
- 160103_I137_FCH3V5YBBXX_L5_WHOSTibkDCAADWAAPEI-74_2.fq.gz
- 160103_I137_FCH3V5YBBXX_L6_WHOSTibkDCAADWAAPEI-74_2.fq.gz
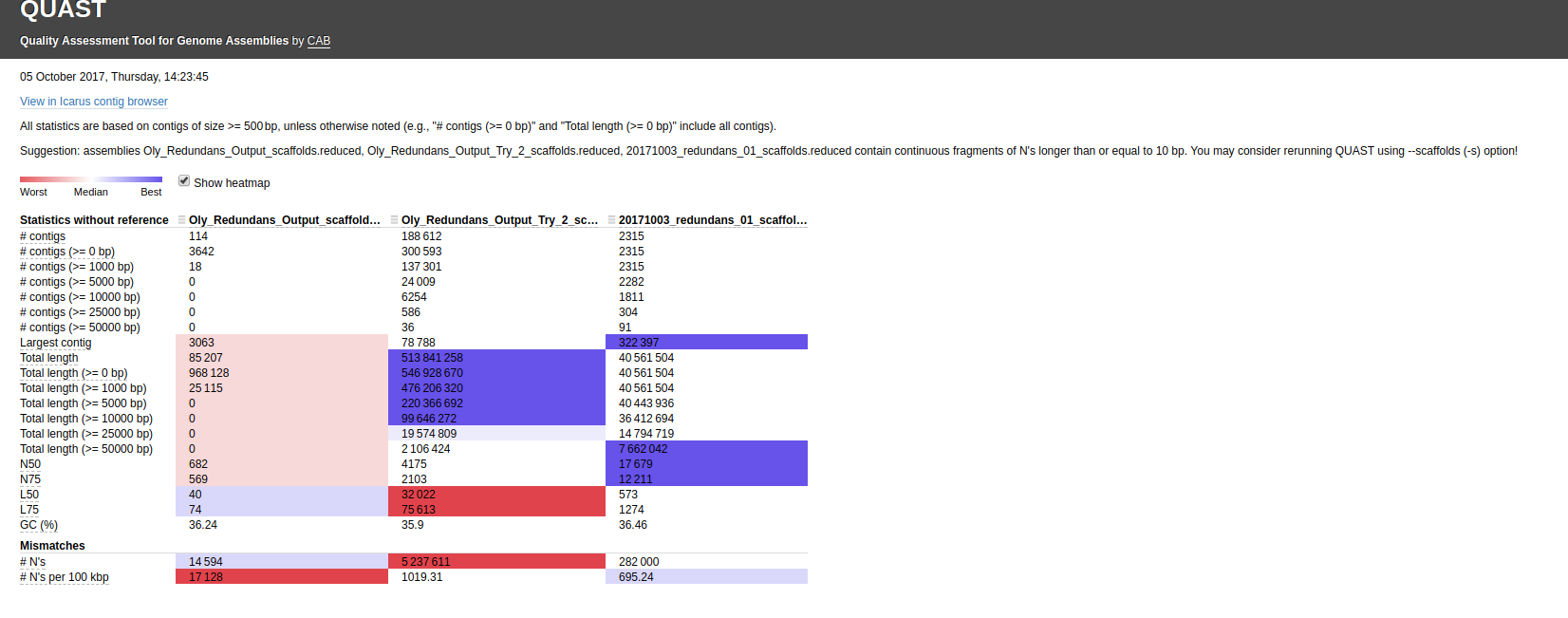
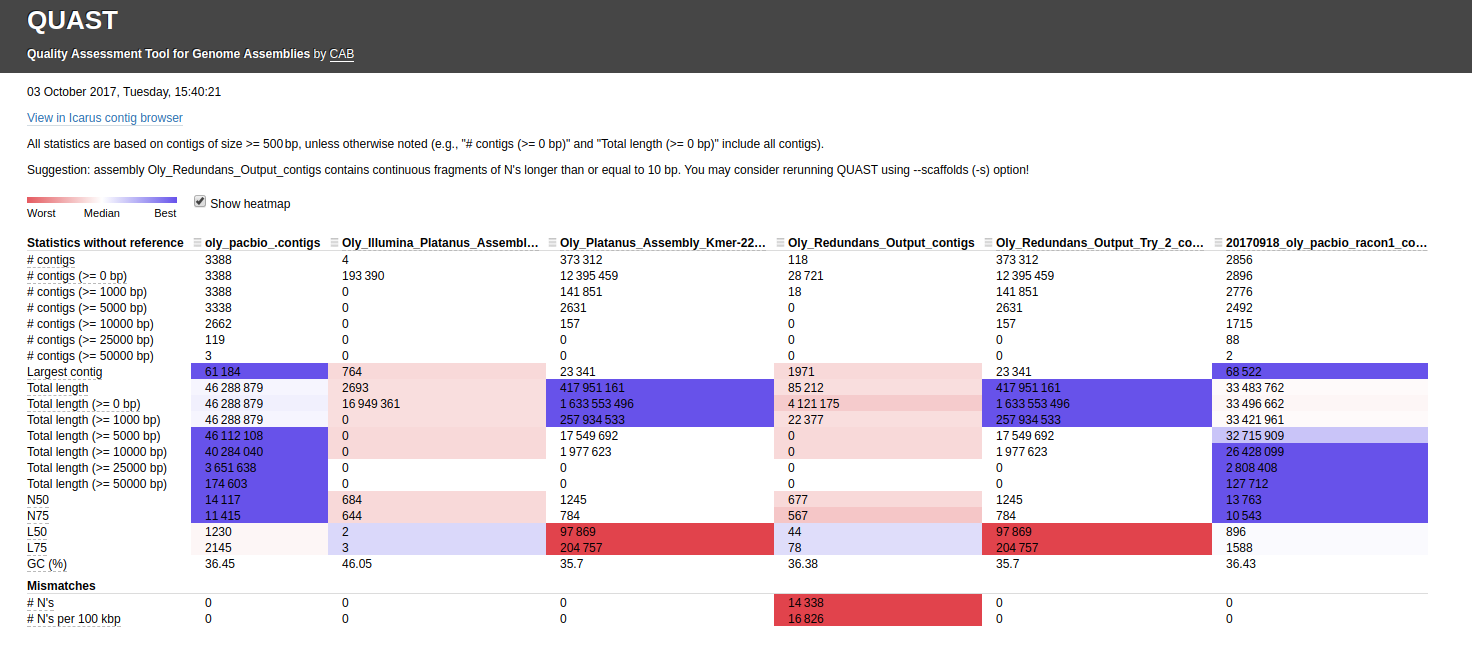
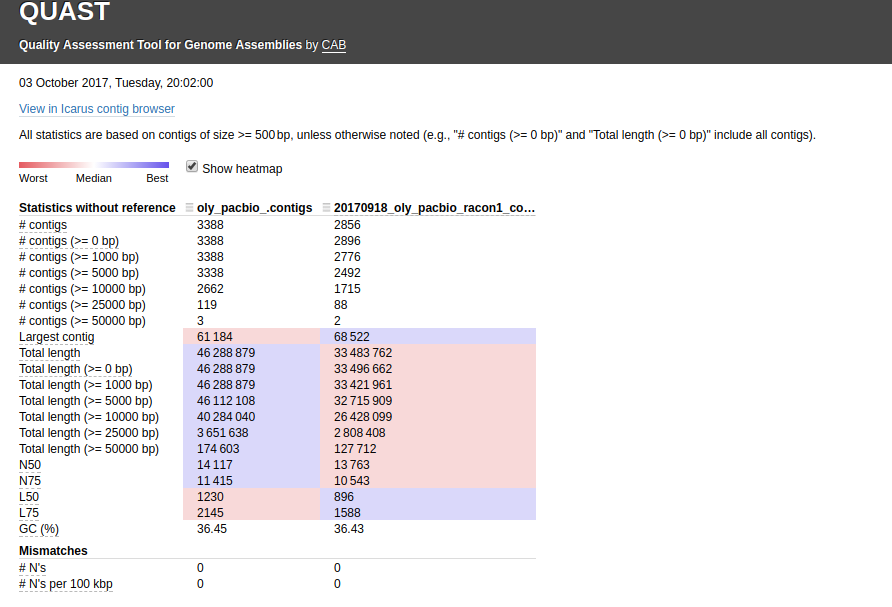
First, I ran QUAST with the default settings:
Interactive link: http://owl.fish.washington.edu/Athaliana/quast_results/results_2017_10_05_14_21_50/report.html
Using that Canu assembly with Redundans certainly seems to results in a better assembly.
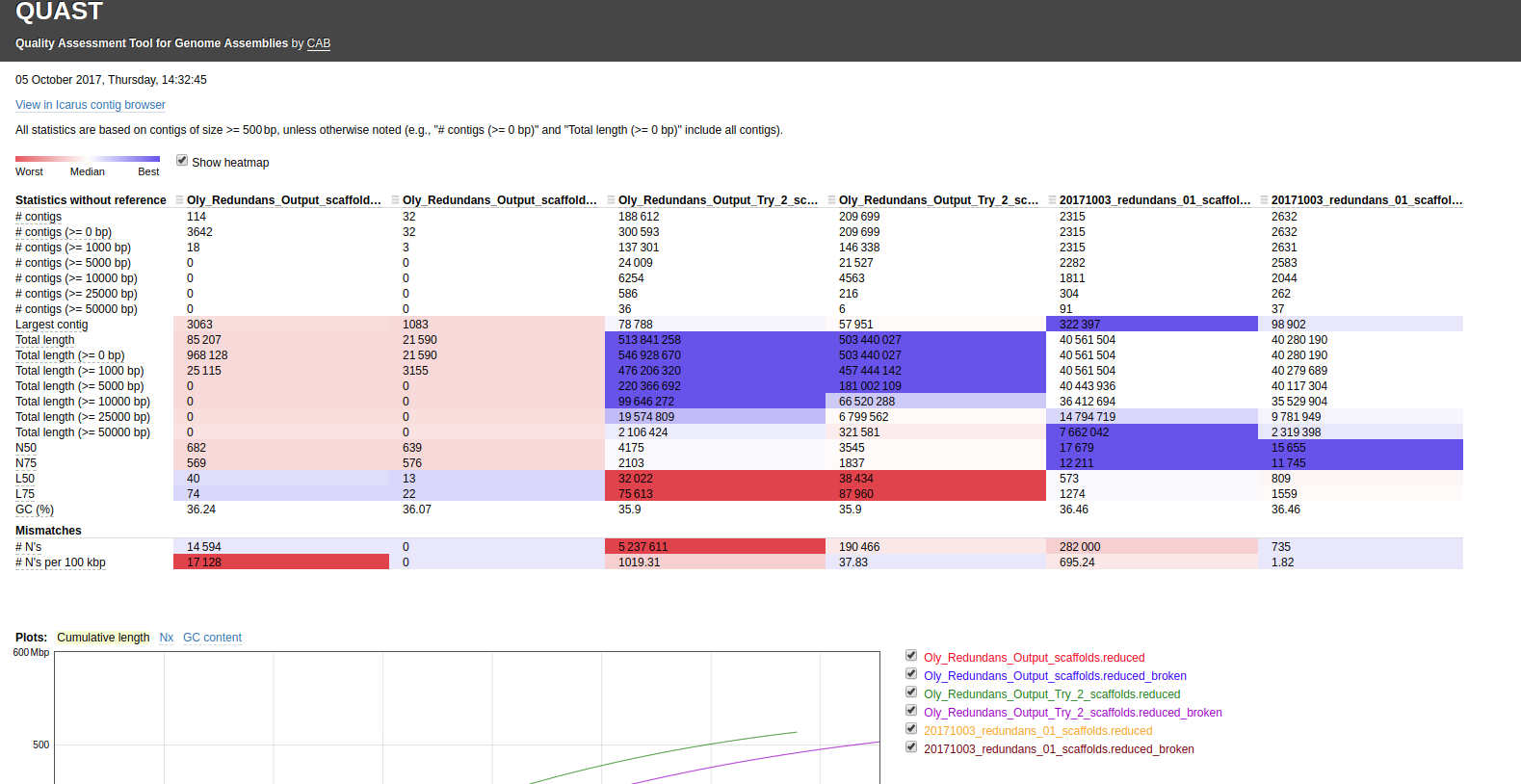
Decided to run QUAST with the –scaffolds option to see what happened:
Interactive link: http://owl.fish.washington.edu/Athaliana/quast_results/results_2017_10_05_14_28_51/report.html
The scaffolds with the “Ns” removed from them are appended with “_broken” – meaning the scaffolds were broken apart into contigs. Things are certainly cleaner when using the --scaffolds option, however, as far as I can tell, QUAST doesn’t actually generate a FASTA file with the “_broken” scaffolds!