
Sample sheet (Google Sheet):
Samples were stored in -80oC:

Sample sheet (Google Sheet):
Samples were stored in -80oC:
Received Caligus tape DNA – two samples:
Female 2 .
Stored in slots H4 and H5 in “Sam’s gDNA Box #2″ in the FTR 213 -20oC freezer.
Google Sheet: Sam’s gDNA Box #2

We received sequencing/assembly data from Phase Genomics.
The data contains two assemblies, produced on two different dates.
All data is here: 20180421_geoduck_hi-c
All FASTQ files (four files; Geoduck_HiC*.gz) were copied to Nightingales:
MD5 checksums were verified and appended to the Nightingales checksum file:
Nightingales sequencing inventory was updated (Google Sheet):
The two assemblies (and assembly stats) they provided are here:
I’ve updated the project-geoduck-genome GitHub wiki with this info.
Received geoduck hatchery metagenome samples from Emma. These samples are intended for DNA isolation.
Admittedly, I’m a bit skeptical that we’ll be able to recover any DNA from these samples, as they had been initially stored as frozen liquid, then thawed, and “supernatant” removed. I’m concerned that the freezing step would result in cell lysis; thus the subsequent removal of “supernatant” would actually be removing the majority of cellular contents that would be released during freezing/lysis.
Here’s the sample prep history, per Emma’s email:
Hi!
Here are the relevant details from my lab notebook:Filters with bacteria to be extracted for proteomics: https://sr320.github.io/Geoduck-larvae-filters/
Each filter was rinsed and cells sonicated:
- Put filter on petri dish on ice
- Use 1-4 mL total to wash front (and back if not obvious where biol material is) of filter while holding with forceps over dish – Use 2 pairs of forceps; I used 4 mL ice cold 50 mM NH4HCO3 to wash inside of filter (filters were folded in half). Washed filters returned to bags and stored at -80C.
- Put wash collected in dish in eppendorf tubes – at this point, remove the amount that will be used for metagenomics (~1/4 of wash) – put 1 mL in metagenome tube (mg) and the remaining was split between 2 tubes for metaproteomics (mp)
These are bacterial cells in ammonium bicarbonate. I spun them down and removed most of the supernatant from each tube.
Let me know if you need any other info!
Box of samples (containing ~38uL of liquid) were stored in FTR209 -20C (top shelf).

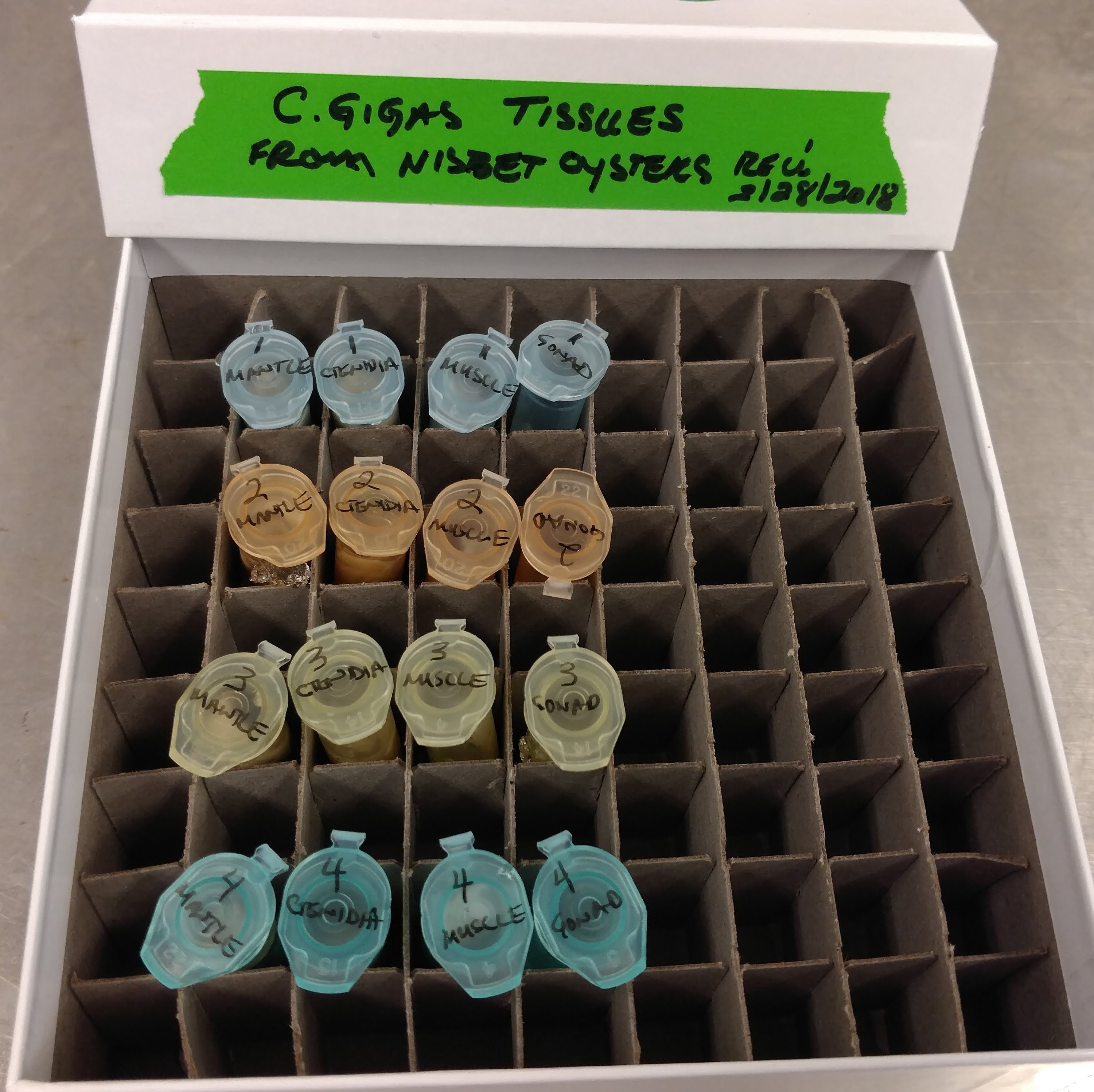
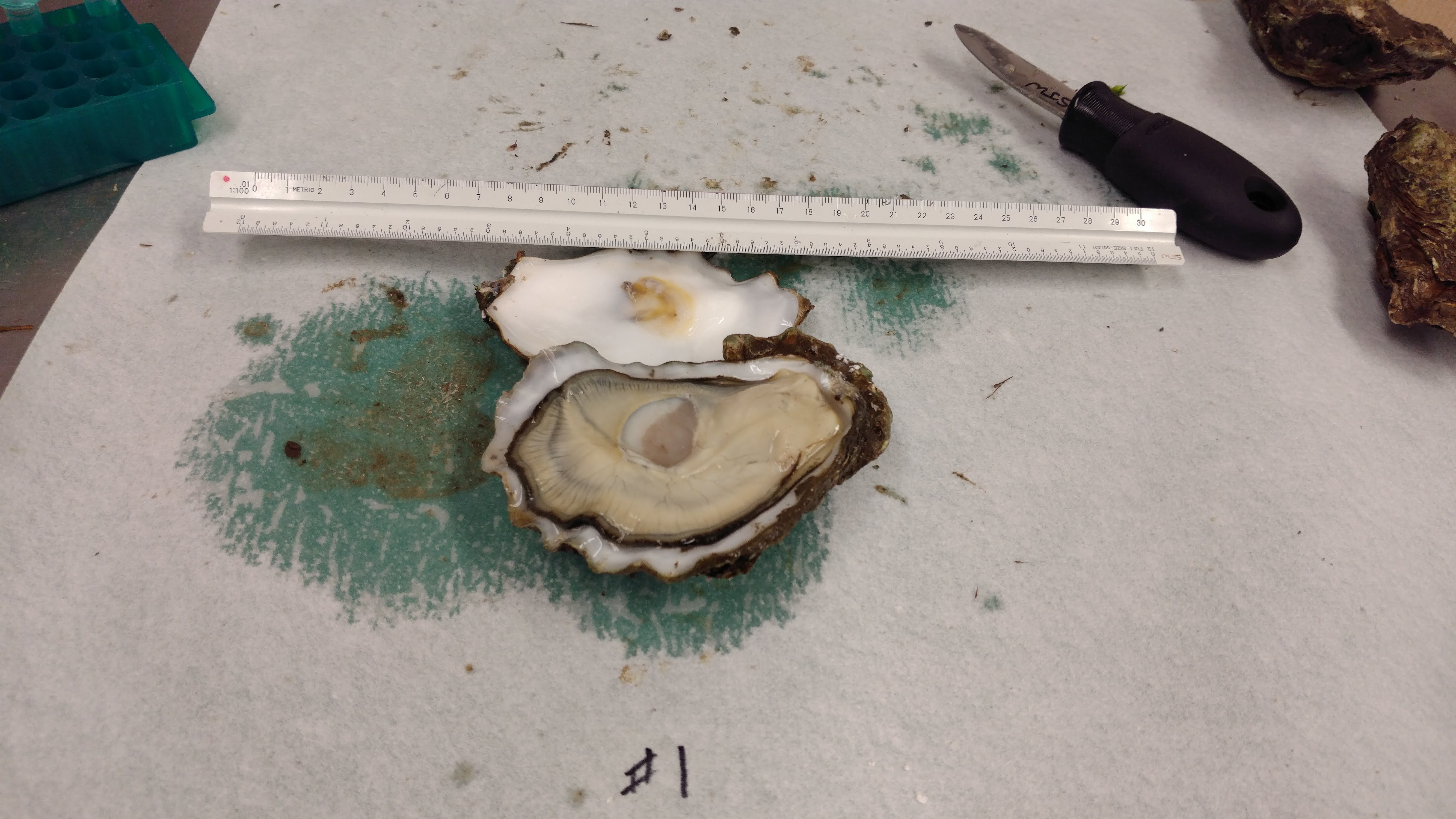
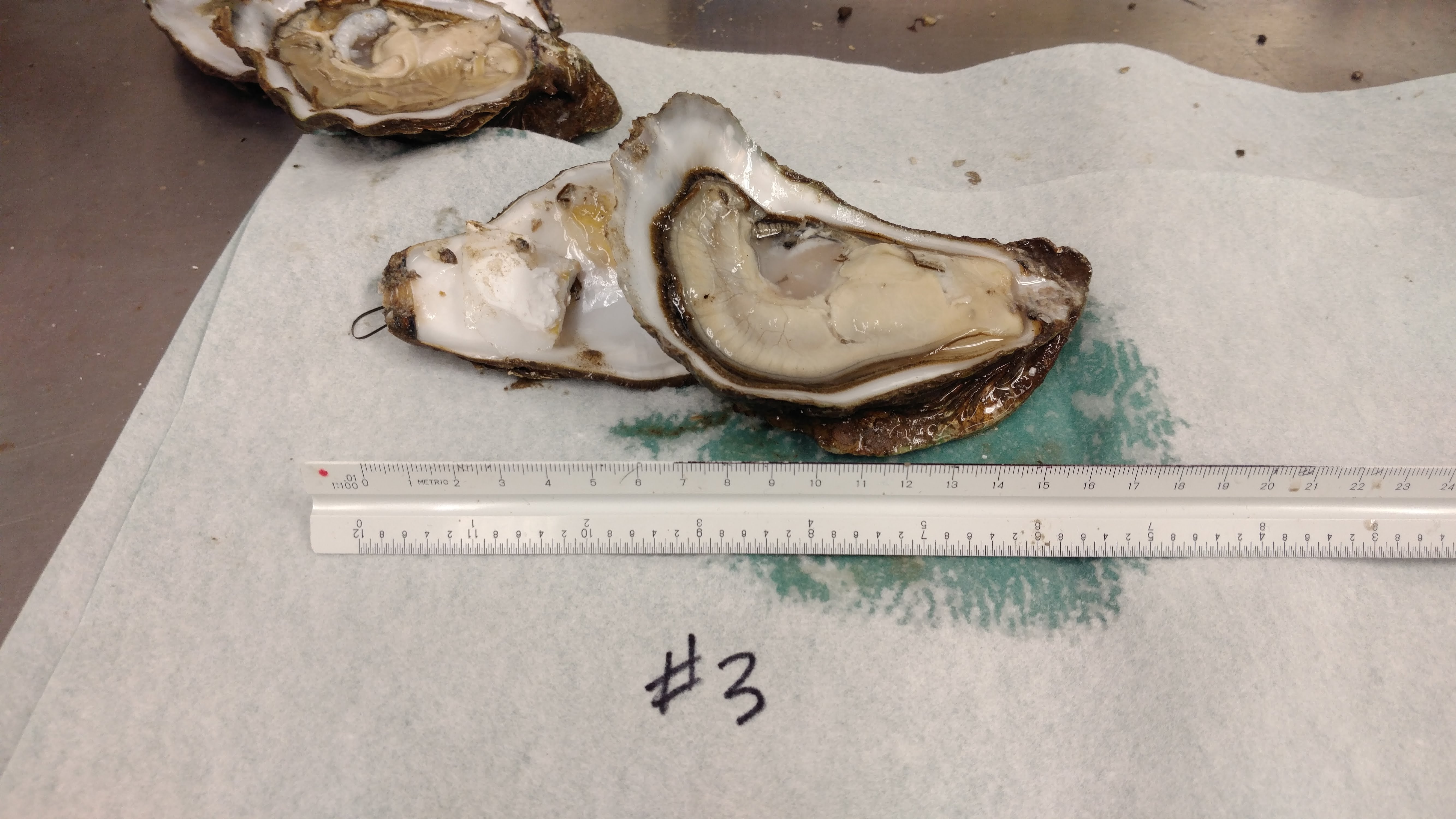
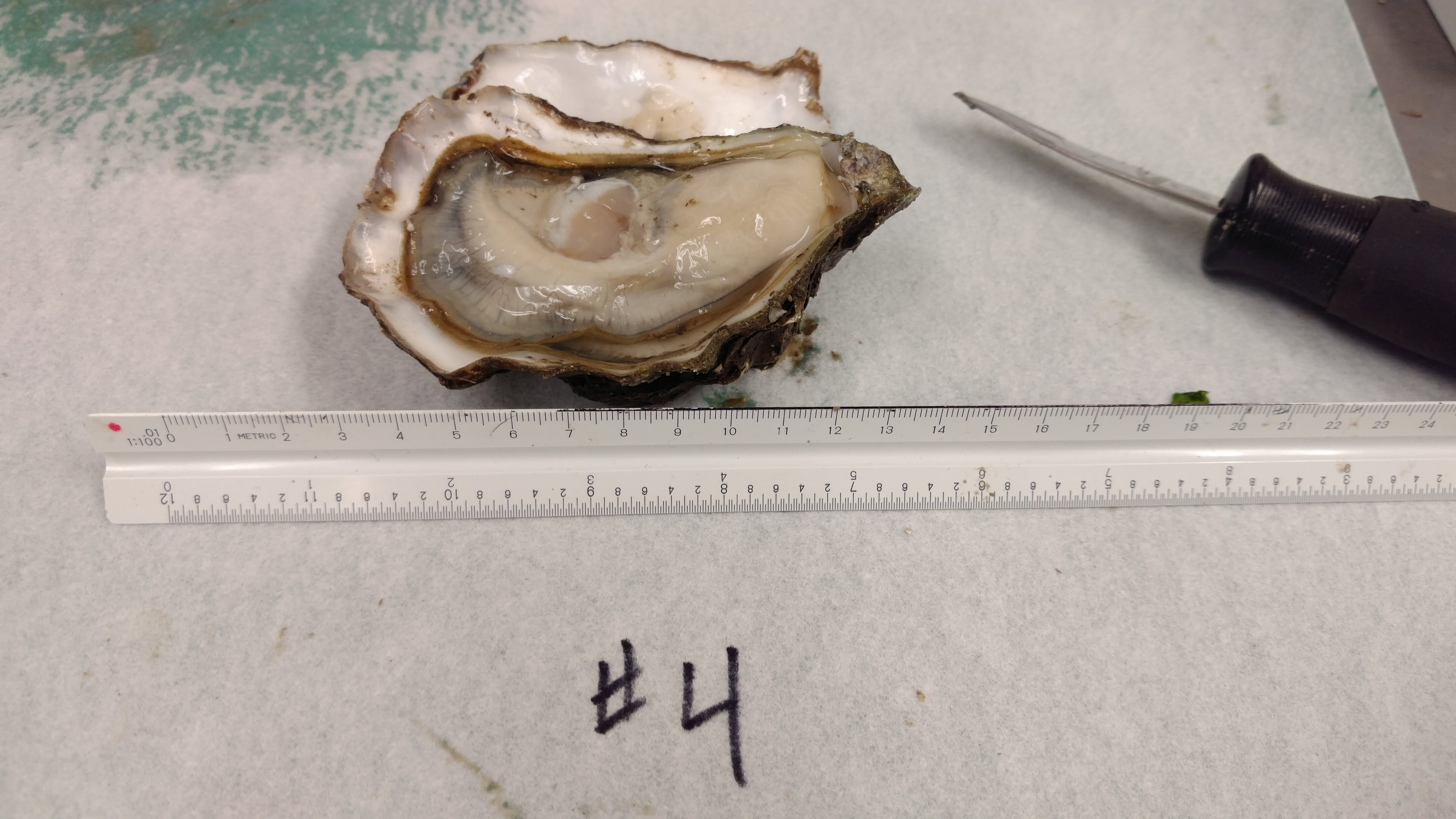
Received a bag of Pacific oysters from Nisbet Oyster Company.
Four oysters were shucked and the following tissues were collected from each:
Utensils were cleaned and sterilized in a 10% bleach solution between oysters.
Tissues were stored briefly on wet ice and then stored at -80C in Rack 2, Column 3, Row 1





Pam Jensen stopped by and dropped off Tanner crab (Chionoecetes bairdi) hemolymph samples (~300) stored in RNA Later (Ambion). Samples were stored at 4C.
Received and stored @-80C in rack 8, row 5, column 5.
The following information was sent with the samples:
| Sample.ID | Date | Temp | pCO2 | Notes |
|---|---|---|---|---|
| 031 | 26-Aug-2016 | 15 | 400 | |
| 032 | 26-Aug-2016 | 15 | 400 | |
| 033 | 26-Aug-2016 | 15 | 400 | |
| 034 | 26-Aug-2016 | 15 | 400 | |
| 035 | 26-Aug-2016 | 15 | 400 | All sample sent; it will be in 2mL screw-cap vial |
| 036 | 26-Aug-2016 | 15 | 400 | |
| 103 | 26-Aug-2016 | 15 | 2800 | |
| 104 | 26-Aug-2016 | 15 | 2800 | |
| 105 | 26-Aug-2016 | 15 | 2800 | |
| 106 | 26-Aug-2016 | 15 | 2800 | All sample sent; it will be in 2mL screw-cap vial |
| 108 | 26-Aug-2016 | 15 | 2800 |
Katie sent this additional info in an email to Steven and me:
These C. virginica samples were exposed to control (400, 6 samples) and OA (2800, 5 samples) conditions for ~4 weeks at 15C. Gonad was carefully extracted by peeling back the outer membrane, flash frozen in liquid N, and placed in -80C (until today when we removed it). During sampling, it was difficult to get a lot of what we considered “pure” gonadal tissue. We sent you ~1/2 of the amount of tissue we have for all samples except for the two samples which were very low and we sent you all the tissue sample we have. Each should be about 10-20 mg of tissue, which I’m worried is not enough for MBD-BS seq. Fingers crossed.
Received histology blocks and slides for Laura Spencer.
Samples were stored in glass cabinet in FTR 213 (see images below).
Received 62 coral (Acropora cervicornis) DNA samples from Javier Casariego at FIU.
Spreadsheet of samples and NanoDrop concentrations provided by Javier (converted to Google Sheet): A.cervicornis_DNA_Extractions(May_2017).xlsx
Samples were temporarily stored at 4c (in FTR 213) until I can perform global methylation assessment on them tomorrow.
Steven received these coral DNA samples today. Here’s his post on Google Plus (stored @ 4C in FTR 213):
Here’s the email from Jose describing the samples:
“Dear Steven, the coral DNA samples were sent today by my student Javier (cc’ed here) to your lab. Here’s an excel attached with info for the samples including concentration and treatment of the coral from which they were extracted (N, nitrogen; NP, nitrogen+phosphorous; C, control).
Please let us know when you get these in the lab so we know all is fine!
thanks!
Chema”
Here’s the spreadsheet he sent (renamed for easier identification later on – original file sent was title DNA Qbit readings), uploaded to Google Drive: